PhD projects
Each Early Stage Researcher has their own well-defined project in Theme 1 or 2, which he/she will implement in research groups under the supervision of members of the Supervisory Board including during their industrial and international secondments. Their technical and transferable skills training has been designed to enhance research effectiveness during their PhDs and prepare them for their post-PhD careers. Suggested previous background for each project is indicated. Others may be appropriate. Supervisors of each project may be found here.
Research Theme 1: Predictive model of bacterial cell division
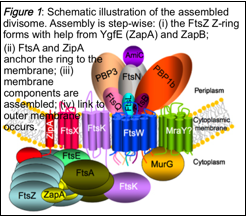
The goal of Research Theme 1 is to develop a molecular-level model for bacterial cell division that can be used to predict how cells will respond to changes in environmental conditions or addition of compounds, e.g. antibacterial agents. The model will be used to pinpoint the key most readily disruptable activities of the division machinery (Figure 1). We shall integrate in vitro experimentation and in vivo testing to determine the minimum number of in vitro experimental parameters required to give accurate predictions of in vivo behaviour, thus reducing the need for live-cell experiments. We believe the time is right to target this challenging goal because the techniques we plan to use have all undergone substantial recent improvements in performance and many of them are poised for the next step forward as indicated below in the 6 projects proposed for this theme. This will enable the required data collection for modelling.
 Research objectives: Cell division involves, at its most basic level, the micron-scale rearrangement (Figure 2) of assemblies of molecules. We shall determine the forces involved, the molecular reactions required to produce the energy to generate the forces, and the structures and arrangements of molecules that achieve this. The end goal is a comprehensive molecular-level predictive model of membrane remodelling during bacterial cell division whose input parameters are determined from biophysics experiments. We shall determine how challenges such as addition of potential drugs and environmental changes will affect the process of cell division without in vivo testing. By integrating a range of techniques from cellular to molecular level we shall develop completely new ways of characterizing molecular processes in cells. The project outputs will include new techniques that will find wide application; new hybrid technologies; integration of theory and experiment in new ways as driven by the clear goal of developing an in vivo model with in vitro input parameters.
Research objectives: Cell division involves, at its most basic level, the micron-scale rearrangement (Figure 2) of assemblies of molecules. We shall determine the forces involved, the molecular reactions required to produce the energy to generate the forces, and the structures and arrangements of molecules that achieve this. The end goal is a comprehensive molecular-level predictive model of membrane remodelling during bacterial cell division whose input parameters are determined from biophysics experiments. We shall determine how challenges such as addition of potential drugs and environmental changes will affect the process of cell division without in vivo testing. By integrating a range of techniques from cellular to molecular level we shall develop completely new ways of characterizing molecular processes in cells. The project outputs will include new techniques that will find wide application; new hybrid technologies; integration of theory and experiment in new ways as driven by the clear goal of developing an in vivo model with in vitro input parameters.
Six coherent individual projects have been designed to achieve the overall goals of the Theme.
The ESR will construct a mesoscopic scale analytical and numerical physical model to describe the responses of the membranes to forces exerted by FtsZ and associated divisome proteins. Combined force and kinetic modelling of the process of cell membrane constriction using data from state-of-the-art EM, optical and force microscopies and spectroscopies will be undertaken. Extension to interactions between the inner and outer membrane will be developed.
The ESR will employ molecular dynamics and course-grain modelling complemented by mass spectrometric (MS) imaging, optical microscopy, and spectroscopic methodologies to identify co-location and orientation of molecular structures and membrane components (lipids and proteins) and determine the atomic-level organisation of molecular complexes in and at membranes on the scale of 10–100 nm with ms time resolution. Recent instrumentation developments of the supervisors make this possible.
This project will focus on overcoming the two major obstacles for producing active membrane protein products: (i) their expression and (ii) their solubilisation in active form. The same issues arise in the biopharmaceutical industry (about half the target receptors are membrane proteins). The ESR will develop novel expression systems to increase output of wild-type and labelled proteins and will develop a new bio-nanotechnology method involving encapsulation of membrane proteins with their immediate lipid environment into discs stabilised by styrene maleic acid.
[]Prokaryotes are subject to stringent selective pressure to optimise their rate of biomass increase in the face of ever-changing ambient conditions. An important environmental factor is the availability of the nutrients and redox substrates required to support the cell's metabolic activities. The cell deploys various kinds of molecular machinery to capture these substrates, transfer these substances into the cell, and process them to channel them into core metabolism. A "Darwinian demon" bacterium would synthesise infinite amounts of each of these types of machinery to extract the environmental resources at infinite rates. In reality, such a demon cannot exist; it is impossible, both physiological and physical reasons. Each piece of molecular machinery represents an investment of building blocks (amino acids) and energy and the cell only has a finite budget at its disposal to make these investments. Thus, the question arises which allocation of investment (i.e. the relative copy numbers of the various types of machinery) gives the best growth yield. This is a formidable problem, made even more challenging by additional factors. For instance, the cells may be able to exploit alternative resources, e.g., a number of different compounds may serve as carbon source. Accordingly, there may be several metabolic modes, which raise several interesting questions: how does the cell effectively choose between these modes, and how does it manage the switch between different modes? This project aims to investigate these fundamental questions, primarily for Gram-negative cells, using a combination of mathematical models, largely based on available data.
The ESR will use leading solid state NMR techniques, linear dichroism (LD) spectroscopy, new Raman spectroscopy methods and molecular modelling to determine the still unknown details of the peptidoglycan structure and determine how the peptidoglycan layer links with the inner membrane and is remodelled into two daughter cells simultaneously with the inner membrane.
In this project the ESR will focus on determining novel ways that cell division can be interrupted. Results from Projects 1.1–1.5 will be integrated to provide understanding of increasingly complex sub-units of the divisome as a function of different perturbations (drugs, environmental conditions) to form a synergistic input.
Research Theme 2: ‘Quality by Design’ (QbD) of pharmaceutical and biopharmaceutical products
There is an increasing push for efficiency in the development and production of pharmaceutical (small molecule) and biopharmaceutical (proteins, nucleic acids, viruses, bacteria) products. The goal of Process Analytical Technology (PAT) is to understand industrial processes by defining their Critical Process Parameters that affect Critical Quality Attributes and monitoring them. Quality by Design (QbD) is increasingly becoming a favoured approach to PAT. The hypothesis underlying QbD for pharmaceutical drugs, is that quality in production can be planned, and that most quality crises and problems relate to the way in which quality was (or was not) planned in the first place. This contrasts with the traditional approaches which depend on Quality Control—effectively, ‘Quality after Design’. The focus of the QbD concept is that quality is built into a product by a thorough understanding of the product itself and the process by which it is developed and manufactured. This understanding is accompanied by an assessment of the risks involved in manufacturing the product and knowledge of how best to mitigate those risks. The European approach to QbD has been developed together with the FDA and Japan and is summarized in International Conference on Harmonisation (ICH) guidelines. However, these developments have made it clear that we do not have the technologies required either for the pre-production phases of QbD, where the goal is to understand the product, nor for the production phases where continuous, fast, non-peturbative in- or on-line testing at the point of manufacture is required.
Theme 2 of CAS-IDP is thus to develop and apply new methodologies to enable QbD to be implemented to develop systems for PAT of pharmaceutical and biopharmaceutical products. The PhD projects (see below) are in both the product-development phase and production phase to enhance product and process robustness. The emphasis will be on hyphenating techniques either to provide complementary information on the same sample or to achieve enhanced specificity or selectivity. We shall focus on hard-to-handle samples including those that: are not very soluble, have high viscosity, lack chromophores for standard techniques, are inhomogeneous, or are at very high concentration. Theme 2 will link with Theme 1 by using Theme 1 molecular systems as test samples for new techniques with the resulting data being inputted to the cell division modelling Theme. This will enhance intra-cohort links between the ESRs and Supervisory Board members as well as making the whole Network intimately aware of how to develop consistent approaches to experimental design and data collection using QbD principles and focussed risk assessments to generate process understanding and establish detailed product/process knowledge.
Most data sets used in Process Analytical Technology (PAT) are reduced to a single parameter to provide a yes or no answer. However, this throws away information about uncertainty that could be used in Quality by Design (QbD). The ESR will take complex data sets and undertake sophisticated statistical analyses (McMC, Bayesian inference and artificial neural networks) to enable complementary data from different techniques (e.g. circular dichroism (CD), NIR spectroscopy and MCD, Projects 2.3 and 2.4) to be combined to give a more complete view of a product. The central aim of the project will be to develop an approach which naturally quantifies uncertainty in parameter estimation, making full use of all available information and can be rapidly applied to experimental data as it emerges. Specific problems that may be considered include the automatic identification of peaks (e.g. in NMR spectra) which are ‘obviously the same’ to the trained human eye but computationally difficult to track; and the effect of uncertainty on the estimation of biochemical reaction networks. The ESR will also undertake selected experiments of their choice with other ESRs to ensure they truly understand issues of experimental data quality and errors (due to the technique and due to the sample). The main emphasis of this project will be on developing and applying new statistical methodologies to data sets produced by the other ESRs.
Potential genotoxic impurities (PGIs) are compounds which can modify DNA and are potentially harmful to human health. In 2007, regulatory guidance was released resulting in PGI detection levels of low parts per million (ppm) relative to the active pharmaceutical ingredient (API) being required. Traditional approaches to detect PGIs employed by industry are time consuming, off-line and labour intensive. The ESR will develop new and novel more sensitive electrochemical detection systems using carbon electrodes that can be integrated into a non-perturbative on-line flow-detection system and do not require chromatographic separation of components. Key objectives are to reduce analysis times, lower costs and improve on current detection levels in the presence of the API. The first targets will be hydrazine and epoxides.
The structure of formulated biopharmaceutical products is currently probed by near and far UV circular dichroism (CD), as required by the European Medicines Agency, in off-line (usually off-site) test facilities, where interference from excipients is often a problem. New challenges are also being brought by the emerging ‘Biosimilars’ market: most simply, what is ‘highly similar’? This emerging market, together with the aims of QbD (which is only beginning to touch biopharmaceuticals), brings urgency to the view that new approaches are required to characterize biopharmaceuticals. In this project the ESR will develop new methods for characterizing biopharmaceutical products uniting the advantages of spectroscopy with circularly polarized light (currently based in the UV region of the spectrum) and the overwhelming importance of near infra red (NIR, 800–2500 nm), NMR, and Raman spectroscopies (incident UV and visible light, output vibrational data) to characterize formulated products. The ESR will determine how to integrate the (single molecule) structural data from ion mobility MS with spectroscopy data that is averaged over the sample and consider the role of data integration from different techniques.
Quantitative electron paramagnetic resonance (EPR) spectroscopy is emerging as a selective tool for identifying radicals and their environment (viscosity, polarity, pH) and biodistribution (by localized measurement or EPR imaging). It is particularly attractive for identifying reactive trace radical impurities resulting from the sterilisation process or to characterize the mechanisms of drug release from biodegradable polymers as it often does not require chromatography before the analysis. In this project the ESR will take the leading EPR innovations developed for carbon-based electronics and develop in-situ EPR methods for pharmaceuticals. The ESR will also pursue the new application of magnetic CD (MCD) to biopharmaceutical concentration determination (based on the distinctive tryptophan signal) and structural characterization. This option has only recently become feasible due to the invention of >1 T small footprint permanent magnets fabricated from alloys of neodymium, iron and boron (NIB). We shall also explore the NIR region and NIR CD spectroscopy.
A mechanistic understanding of active pharmaceutical ingredient dissolution from its formulation matrix is essential for understanding a drug’s pharmacokinetic profile and optimising performance of active ingredients immobilised in the matrix. This is challenging for many drug formulations but particularly so for lipid-based drug delivery systems where different batches of lipid-coated polymers vary based on the source of the lipids and the polymerization methods used to create them. This variability leads to unpredictable stability issues with the drugs. To address these challenges the ESR will implement state-of-the-art sophisticated solid-state nuclear magnetic resonance (NMR) methodologies to identify and quantify matrices used in formulations as a function of time. Ss-NMR will be complemented by solution NMR, mass spectrometric, New NIR, and Raman methods in collaboration with Project 2.3.
With biopharmaceuticals one has to confirm the molecular identity of the active ingredient, ensure it is consistently formulated with a well-defined oligomeric state, and is folded into the correct molecular structure (cf. Project 2.3). However, at our current state of expertise, this is not sufficient to ensure consistent batch-batch bioactivity. For this purpose assays such as enzyme-linked immunosorbent assays (ELISA) or complement deposition assays are employed. However, such assays are challenging to perform and not amenable to an instant read-out on-line format. In this project the ESR will develop the use of LD spectroscopy as an on-line detector for biophamaceutical activity by measuring binding of active ingredients to antibodies which are themselves covalently attached to flow-oriented bacteriophage.
A mechanistic understanding of active pharmaceutical ingredient dissolution from its formulation matrix is essential for understanding a drug’s pharmacokinetic profile and optimising performance of active ingredients immobilised in the matrix. Two significant problems with the traditional dissolution reaction chambers approach are: (1) the hydrodynamics of the stirring are neither controlled nor understood, therefore separating mass transport effects from dissolution/release kinetics is extremely difficult; (2) information is only provided on bulk concentration changes not on what is happening at the drug-matrix surface. The ESR will radicalise the way dissolution processes are studied in the pharmaceutical industry by developing miniaturised flow cells, where the hydrodynamics of the drug dissolution process is controlled and then modelled as a function of biologically relevant variables (temperature, pH, flow rate, hydrophobicity, and viscosity). Spectroscopic and electrochemical techniques will be used to study concentration and morphology changes at the surfaces of tablets and microcrystalline formulations. Solid-state NMR will also be used to determine the solid state components as a function of time.
