Evidence of Salmonella Paratyphi C found for the first time in medieval northern Europe
• Eight hundred year old Norwegian skeleton found to have traces of Salmonella
• Reshapes understanding of the bacterial pathogen Salmonella enterica
• EnteroBase genomic database used to identify genomes
• Possible link to domestication of pigs
Genome research conducted by the University of Warwick suggests that enteric fever, a potentially lethal disease more commonly found in hot countries, was present in medieval Europe.
Salmonella Paratyphi C causes enteric fever, a life-threatening infection, and has been detected in a 800 year old human skeleton discovered in Trondheim, Norway. Now scientists are speculating that the evolution of enteric fever could be linked to the domestication of pigs across northern Europe.
The research was conducted by a team of international collaborators led by Professor Mark Achtman of the University’s Warwick Medical School and their paper Pan-genome Analysis of Ancient and Modern Salmonella enterica Demonstrates Genomic Stability of the Invasive Para C Lineage for Millennia has been published in the journal Current Biology.
He and his team analysed bacterial DNA found in the teeth and bones of the skeleton of a young woman who is believed to have migrated to Trondheim from the northernmost areas of Scandinavia or Northwest Russia by her early teens only to die there around the age of 19-24 years.

They reconstructed a genome of Salmonella Paratyphi C which causes enteric fever in areas of poor sanitation and lack of clean drinking water. Their discovery indicates that the young Norwegian died of this disease and suggests that these bacteria have long caused enteric fever across northern Europe.
Prof essorAchtman said: “Paratyphi C is very rare today in Europe and North America except for occasional travellers from South and East Asia or Africa, where the disease is more common.
"This is the first time that any Salmonella have been found in old human remains in Europe, which is surprising because other Salmonella are more common today, including Salmonella causing typhoid fever, called Typhi, and Salmonella causing food poisoning. Earlier this year, Vågene and co-authors described related Paratyphi C from skeletons in Mexico, who died in 1545 CE, and speculated the Paratyphi C entered the Americas together with Europeans.”
The new results included comparative analyses of the Paratyphi C genome found in the skeleton against modern Salmonella genome sequences from EnteroBase, an online database developed at the University of Warwick and used internationally.
This revealed that Paratyphi C represents the evolutionary descendants of a common ancestor, or clade, within the Para C lineage. The Para C Lineage includes Choleraesuis, which causes septicaemia in pigs and boar and Typhisuis which causes epidemic swine salmonellosis (chronic paratyphoid) in domestic pigs. These different host specificities likely evolved in Europe over the last 4,000 years and coincide with the timing of pig domestication in Europe.
According to historical records, humans have long been afflicted by bacterial infections, yet genomic analyses of living bacterial pathogens routinely estimate a date for the most recent common ancestor of no more than a few centuries. In general, evolutionary trees contain a stem group, which may include lineages that are now rare or extinct, as well as the crown group of living organisms.
Historical reconstructions based only on the crown group ignore the older sub-lineages in the stem group and thereby provide an incomplete picture of the older evolutionary history of the pathogen. In contrast, analyses of ancient DNA such as the Paratyphi C genome can shed light on additional millennia of bacterial pathogen evolution that occurred prior to the origin of the crown group.
Professor Achtman added: “Using EnteroBase we were able to define the Para C lineage from 50,000 modern Salmonella enterica genomes and find that over its 3,000 year history only a few genomic changes occurred within the Para C lineage.
“As well as reshaping our understanding of Salmonella enterica, our research has triggered intriguing speculations about historical host jumps during the Neolithic period between humans and their domesticated animals.”
19 July 2018
Photocaptions:
photo 1 The face reconstruction of the young woman (credit: Caroline Wilkinson, Mark Roughley, Ching Liu and Kathryn Smith at the Face Lab at Liverpool John Moores University for their different contributions to the production of the facial depiction)
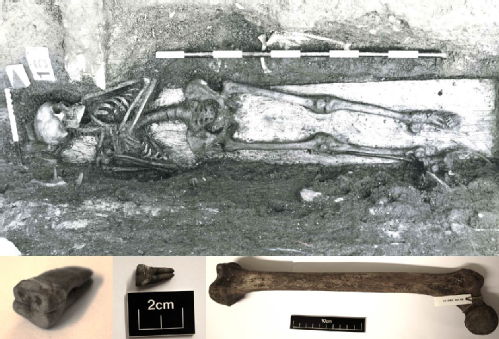
photo 2 The skeleton plus teeth and bones from skeleton
Notes to editors
Pan-genome Analysis of Ancient and Modern Salmonella enterica Demonstrates Genomic Stability of the Invasive Para C Lineage for Millennia is published in Current Biology.
DOI https://doi.org/10.1016/j.cub.2018.05.058
Authors
Zhemin Zhou, Warwick Medical School, University of Warwick, Gibbet Hill Road, Coventry CV4 7AL, UK
Inge Lundstrøm, Centre for GeoGenetics, Natural History Museum of Denmark, University of Copenhagen, Øster Voldgade 5-7, 1350 Copenhagen, Denmark
Alicia Tran-Dien, Unité des Bactéries Pathogènes Entériques, Institut Pasteur, Paris, France
Sebastián Duchêne, Department of Biochemistry and Molecular Biology, University of Melbourne, Parkville, Victoria 3010, Australia
Nabil-Fareed Alikhan, Warwick Medical School, University of Warwick, Gibbet Hill Road, Coventry CV4 7AL, UK
Martin J. Sergeant, Warwick Medical School, University of Warwick, Gibbet Hill Road, Coventry CV4 7AL, UK
Gemma Langridge,Wellcome Trust Sanger Institute, Cambridge, UK (Present address: Molecular Microbiology, Norwich Medical School, University of East Anglia, Norwich, UK)
Anna K. Fotakis, Centre for GeoGenetics, Natural History Museum of Denmark, University of Copenhagen, Øster Voldgade 5-7, 1350 Copenhagen, Denmark
Satheesh Nair,Wellcome Trust Sanger Institute, Cambridge, UK (Present address: Gastrointestinal Bacteria Reference Unit, Public Health England, London, UK)
Hans K. Stenøien, NTNU University Museum, N-7491 Trondheim, Norway
Stian S. Hamre, Department of Archaeology, History, Cultural Studies and Religion, University of Bergen, Post Box 7805, 5020 Bergen, Norway
Sherwood Casjens, Pathology Department, University of Utah School of Medicine, Salt Lake City, UT 84112, USA
Axel Christophersen, NTNU University Museum, N-7491 Trondheim, Norway
Christopher Quince, Warwick Medical School, University of Warwick, Gibbet Hill Road, Coventry CV4 7AL, UK
Nicholas R. Thomson,Unité des Bactéries Pathogènes Entériques, Institut Pasteur, Paris, France
François-Xavier Weill; Wellcome Trust Sanger Institute, Cambridge, UK
Simon Y. W. Ho, School of Life and Environmental Sciences; University of Sydney, Sydney NSW 2006, Australia
M. Thomas P. Gilbert ,Centre for GeoGenetics, Natural History Museum of Denmark, University of Copenhagen, Øster Voldgade 5-7, 1350 Copenhagen, Denmark; NTNU University Museum, N-7491 Trondheim, Norway
Mark Achtman, Warwick Medical School, University of Warwick, Gibbet Hill Road, Coventry CV4 7AL, UK
For more information contact Nicola Jones, Media Relations Manager, University of Warwick N.Jones.1@warwick.ac.uk or 07920531221
