Metallo Foldamer-Proteins Hybrids: Novel Biomimetic Scaffolds for Sensing and Imaging Applications
Principal Supervisor: Dr Sarah Pike
Secondary Supervisor(s): Dr Anna Peacock
University of Registration: University of Birmingham
BBSRC Research Themes: Understanding the Rules of Life (Structural Biology, Systems Biology)
No longer accepting applications
Project Outline
Foldamers are synthetic helical oligomers that adopt stable secondary structures through mimicking the folding patterns of biological systems to generate structures of well-defined size and shape (Figure 1).1 Foldamers have been the subject of great interest due to their diverse range of applications in supramolecular chemistry. However, despite the importance of biomimetic foldamers and their known ability to mimic the simple natural helical topologies, there are no reports on the development of higher order foldamer scaffolds that can mimic the sophisticated tertiary and quaternary structures found in biological systems. Given that it is often this higher structure that controls the properties and, in turn the function, of biological structures, accessing new higher order foldamer topologies has the exciting potential to open up the field towards the design of truly artificial biological structures. Such structures would hybridise the advantages of biology with those of chemistry, for potential applications in catalysis or sensing, to name but a few.
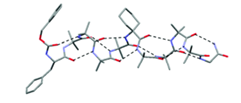
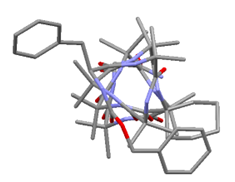
Figure 1. Stable folded conformation of synthetic organic helical oligomer (foldamer) in the solid state. a) viewed side on, b) viewed from the N terminus
Miniature artificial protein structures already exist in the form of de novo designed assemblies. The majority of these are based on the coiled coil, a supercoiling of a defined number (two or more) of alpha-helices, held together through supramolecular interactions. These scaffolds can further be stabilised by the introduction of a metal ion binding site within the core of the coiled coil thereby linking the individual helixes. In recent years, de novo metallopeptide coiled-coil scaffolds, have found impressive applications beyond the repertoire of what Biology has evolved to do, such as MRI contrast agents.3 However, the fact that these existing metallopeptide coiled-coil scaffolds are made from biological building blocks, means that they are based on, and therefore limited, to biological shapes and structures. Accordingly, existing systems are restricted to exhibiting a range of shapes and structures, that historically have been limited by evolutionary-imposed constraints, and this severely hinders the optimisation of their performance and the accessibility of new scaffolds for innovative applications.
To truly realise the full potential of synthetic biology it is vital that we are able to access new topologies that are not restricted to biomimetic systems with evolution-imposed constraints. Accordingly, we will create new hybrid metallofoldamer-protein systems that adopt higher order coiled coil assemblies which are able to adopt a range of novel topologies for generating new applications as potential sensors for biological analytes or effective medical diagnostics tools.
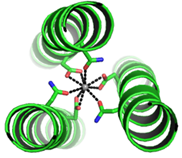
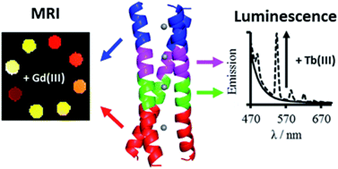
Figure 2. a) Coiled-coil scaffold incorporating lanthanide ions, b) de novo coiled-coil lanthanide design for potential MRI imaging and luminescence purposes
Objectives
- To create a new class of artificial protein scaffold, based on metallofoldamer scaffolds inspired by metallo coiled-coils, and fully characterize them using standard and advanced analytical and spectroscopic techniques.
- To generate libraries of metallofoldamer coiled coil scaffolds wherein fundamental structural features (e.g., type of metal) are systematically varied to generate a library of hybrid scaffolds
- To interrogate the designed library and to establish “sequence-structure-function” relationships on the influence of key structural features (e.g., foldamer length, foldamer building block and/or type of metal) on their performance as sensors for biological analytes and as luminescent or magnetic probes for medical imaging applications.
Methodology
Each of the objectives of the project require a combination of organic chemistry synthesis and advanced analytical study to generate the new artificial protein, metallofoldamer-coiled coil, scaffolds and to assess their stability and performance as sensors and as luminescent or magnetic probes. This work will be carried out under the supervision of Drs. Pike and Peacock in the Department of Chemistry.
References
- G. Guichard, I. Huc, Chem. Commun., 2011, 47, 5933-5941.
- H. R. Marsden, A. Kros, Angew. Chem. Int. Ed., 2010, 49, 2988-3005.
- A. F. A. Peacock et al., Dalton Trans., 2018, 47, 10784.
Techniques
- Cell culture
- Super-resolution fluorescence microscopy
- Data analysis (R, Matlab, Python or C depending on the student preference)
- Bioinformatics, data curation and interaction with public databases