Zak Ogi-Gittins
About me & Research Interests
Hello! I am a 3rd year PhD student at the Mathematics for Real-World Systems Centre for Doctoral Training. Since joining the MathSys CDT, I have become interested in Mathematical Epidemiology, especially given the attention that the field has gained during the COVID-19 pandemic. I am also interested in applied dynamical systems and parameter inference.
Publications and References
[1]: A simulation-based approach for estimating the time-dependent reproduction number from temporally aggregated disease incidence time series data, I. Ogi-Gittins, W.S. Hart, J. Song, R.K. Nash, J. Polonsky, A. Cori, E.M. Hill, R.N. Thompson, Epidemics, Volume 47, 2024, 100773.
Academic Projects
MathSys Group Project: "Adaptive management during an ongoing pandemic"
I worked within a small group of MathSys students to evaluate the impact of using Adaptive Management techniques to improve decision making during an epidemic. The logic underpinning Adaptive Management is that parameter uncertainty is likely to be resolved as an outbreak unfolds. Mathematically, we can use the information that our parameter knowledge will change in the future to aid our decision-making processes today. Ideally, these strategies would be used in present-day epidemic modelling but significant work is required for this approach to be more popular in the modelling community.
In our project, we attempted to evaluate the optimum time that the UK should have begun lockdown (given the up to date knowledge at the time) using Adaptive Management methodology. We found (as is well documented in the literature) that the decision-making process is highly dependent on the loss function, i.e. a linear combination of lives lost and time spent in lockdown.
MathSys Masters Dissertation: "How mis-specifying time-varying generation intervals affects reproduction number inference"
Statistical monitoring of epidemics relies on the data streams that are provided. These streams suffer multiple biases, i.e. delay-times, under-reporting, etc. In spite of these biases, there are several methods that can eliminate some of the issues when inferring the time-dependent reproduction number during an ongoing epidemic [Cori et al, Thompson et al., Epi-Estim]. In this report, we investigated the effects of generation-interval mis-specification on reproduction number inference, and which properties of misspecification still enable the user to process whether the inference is likely to be an over/under-estimate.
Highly Pathogenic Avian Influenza Rapid Response
During the UK’s H5N1 (bird-flu) outbreak, a small group within SBIDER (Systems Biology & Infectious Disease Epidemiology Research) began modelling the spread between poultry holdings across England, Wales and Scotland. This piece of work uses a spatial model established by Keeling et al., which enables a significant improvement in the speed of computation.
From January-March 2022, we conducted an initial analysis using MCMC algorithms to estimate the dispersion kernel present in the model, as well as other key parameters such as transmissibility and susceptibility parameters. From this, we were able to simulate counterfactual scenarios to evaluate the relative risk of the outbreak that occurred in North Yorkshire. We also modelled the impact of ring-culling and enhanced surveillance.
Since then, I have worked on extending this initial project to look at different models, including a spark-term model, which accounts for wild bird incursions (see posteriors for fitted parameters below). We plan to compare the models that we have fitted and continue our counterfactuals analysis.
Inference of the time-dependent reproduction number from temporally aggregated incidence data
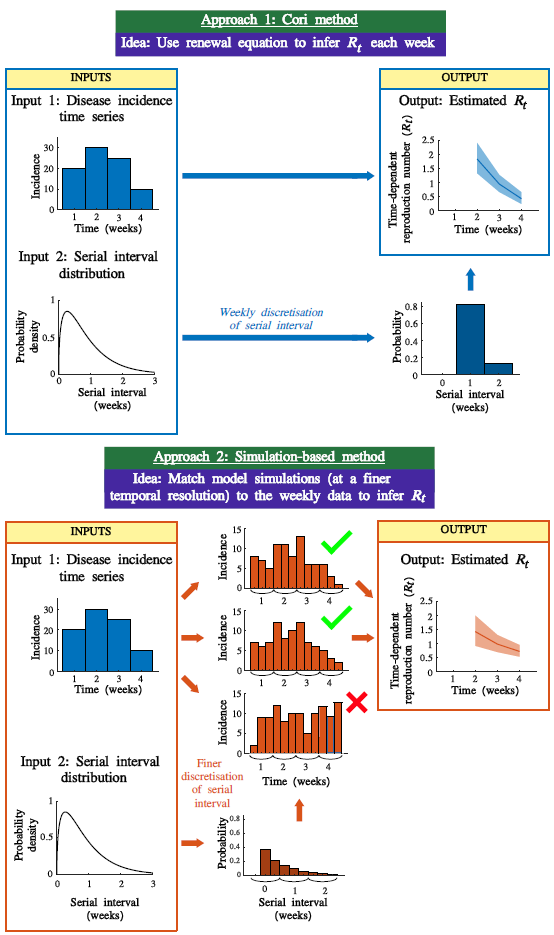
One of the first statistics epidemiologists want to estimate during an outbreak is the time-dependent reproduction number since this enables us (scientists, policy-makers, the public) to gain a crude but very informative understanding of the scale of transmission from the average infectious person. A problem that besets reproduction number inference is that incidence data is often collected on a time-scale that is shorter than a typical generation interval (the time between infections in an infector-infectee pair). This implies that, with a high probability, infectious individuals recording their infection date at time-step t were infected by another individual recording their infection on the same time-step. This is hugely consequential to Rt inference because the renewal equation that models these inference processes assumes that infections on day t arise due to due cases prior to day t.
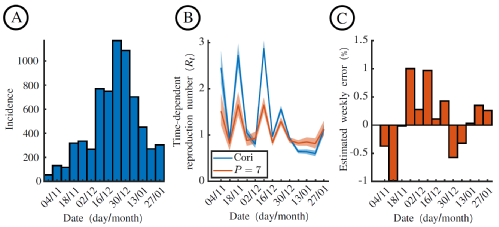
In June 2024, we published a paper in Epidemics [1], outlining how we can use a simulation based approach to infer Rt by simulating the epidemic on a shorter time-scale and then matching ('Exact Approximate Bayesian Computation') the shorter time-step simulated incidence to the incidence data. The following figures are the schematic illustrating the approach, and example inference panels for how distinct the resulting Rt inference (between the standard Epi-Estim/Cori approach and this novel approach) is when applied to an outbreak of ILIs (Influenza Like Illnesses) in Wales (2019-20).
Inference of the time-dependent reproduction number from temporally aggregated and under-reported incidence data
Our colleague in Oxford (Nic Styen) and my supervisor (Robin Thompson) extended the existing method (see above section) to account for under-reporting as well as temporally aggregated incidence data. Not only does this method account for an additional source of uncertainty, it actually speeds up computation time. This is because we are now able to drop the 'Exact ABC' approach and instead accept all simulated weeks that exceed the reported incidence on that week. The key insight is that some simulated weeks are more likely than others, and therefore we weight simulations proportional to their likelihood.
This latter work is currently unpublished but we presented the ideas and results at the SMB conference in Seoul, South Korea in July 2024. We hope to publish this work in due course.

Teaching Responsibilities
In my first year, I was a TA on the MA398 and MA4M1 modules in the Warwick Maths Department, and in my second year, I continued TAing on the MA4M1 module.
Contact Information
Office: D2.11 Complexity Science
Zeeman Building,
University of Warwick,
Coventry,
CV4 7AL