Research
Research in the Jenner Lab focuses on the application of mass spectrometryLink opens in a new window, in combination with other analyticalLink opens in a new window and structural techniquesLink opens in a new window, to solve complex biological problems.
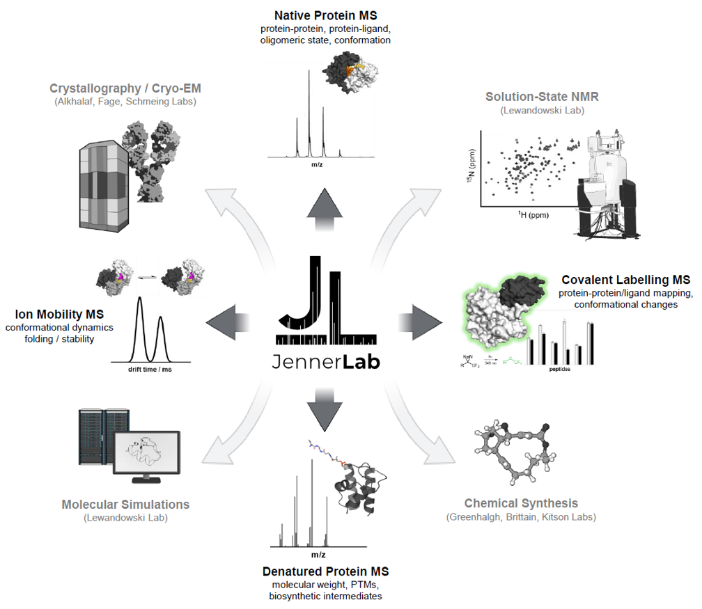
Our primary interest concerns the enzymes involved in the biosynthesis of polyketideLink opens in a new window and non-ribosomal peptideLink opens in a new window natural products – this includes a fundamental understanding of their structures, mechanisms and protein-protein interactions. Our key research themes are outlined below.
Enzymology and Interactions of Megasynth(et)ases.
Many important pharmaceuticals and agrichemicals are derived from polyketideLink opens in a new window and non-ribosomal peptideLink opens in a new window natural products. These complex and diverse classes of molecules are biosynthesised by polyketide synthasesLink opens in a new window (PKSs) and non-ribosomal peptide synthetasesLink opens in a new window (NRPSs), respectively. Often referred to as ‘megasynth(et)ases’, both PKS and NRPS systems are enormous multidomain enzymatic complexes that function in a highly programmed manner to achieve biosynthetic fidelity. Central to many biosynthetic systems is a small (~10 kDa) carrier protein domainLink opens in a new window that is post-translationally modified with a 4'-phosphopantetheine prosthetic ‘arm’, to which intermediates and extender units are covalently tethered via a thioester bond. The carrier protein sits at the heart of multi-domain enzymes such as PKSs and NRPSs, engaging in specific protein interactions with each catalytic domain during the biosynthesis. Our interests focus on understanding the structures, interactions and mechanisms of these systems from various microbial origins.
Representative Publications: Chem. Sci., 2025, 16, 13173–13182; JACS Au, 2025, 5, 144–157, Nat. Commun., 2023, 14, e2832; Nat. Chem. Biol., 2022, 18, 1410–1416; Nat. Commun., 2022, 13, e62; Chem. Sci., 2021, 12, 13676–13685; Chem. Sci., 2020, 11, 11525–11530; Nat. Chem. Biol., 2018, 14, 270–275.
Genomics-Driven Natural Product Discovery & Biosynthetic Pathway Elucidation.
Bacteria and fungi are prolific producers of structurally complex, highly bioactive natural products. Advances in post-genomic techniques and the propensity for biosynthetic genes to group together – termed biosynthetic gene clusters (BGCs) - have allowed the prediction of pathways from genetic sequences alone and have facilitated numerous targeted discovery methods. In this area, our current focus is on Gram negative bacteria (BurkholderiaLink opens in a new window and PseudomonasLink opens in a new window), fungi of the AscomycetesLink opens in a new window genus, and the development of effective genomics-driven natural product discovery platforms to exploit these organisms.
Representative Publications: mBio, 2021, 12, e00715–00721; Angew. Chem. Int. Ed., 2020, 59, 23145–23153; Chem. Sci., 2019, 10, 5489–5494; Nat. Microbiol., 2019, 4, 996–1005.

