Lewandowski Group
Lewandowski GroupLewandowski group focuses on development and applications of solid-state and solution NMR to study structure, dynamics and interactions of biological systems in the context of integrated structural biology. We are associated with the Warwick solid-state NMR grouping and Chemical Biology Research Facility. Research Themes | NMR Facilities | Available Positions
Solid-State NMR MethodologyWe develop methods to probe structure and dynamics of molecules. Protein structure & interactionsWe apply both solid- and solution-state NMR to various biomolecular systems. Natural products biosynthesisStructural biology and engineering of natural products biosynthetic machinery. Protein dynamicsExperimental determination and simulation of molecular motion. AntibioticsUnderstanding modes of action and facilitating designing new antibiotics. |
News
|
 |
Solid-State NMR Methodology DevelopmentWe develop solid-state NMR methods for studying structures and dynamics of proteins. Highlights:
|
 |
Protein structure determination and molecular interactionsWe employ a combination of different structural biology and biophysical methods led by solid-state and solution-state NMR to investigate structure, dynamics and interactions of various biological systems. We work on systems that are natural targets for solid-state NMR, e.g. nanocrystalline proteins, protein complexes (e.g. protein-antibody complexes and complexes from polyketide synthases), fibrils and membrane proteins and systems that benefit from combined solution/solid state NMR approaches. We are also interested in characterization of molecular interactions, in particular protein interactions with other proteins, ligands and nucleic acids. Main areas:
|
 |
Protein dynamicsWe apply a variety of methods centered around NMR to characterize protein motions at atomic resolution. Highlights:
|
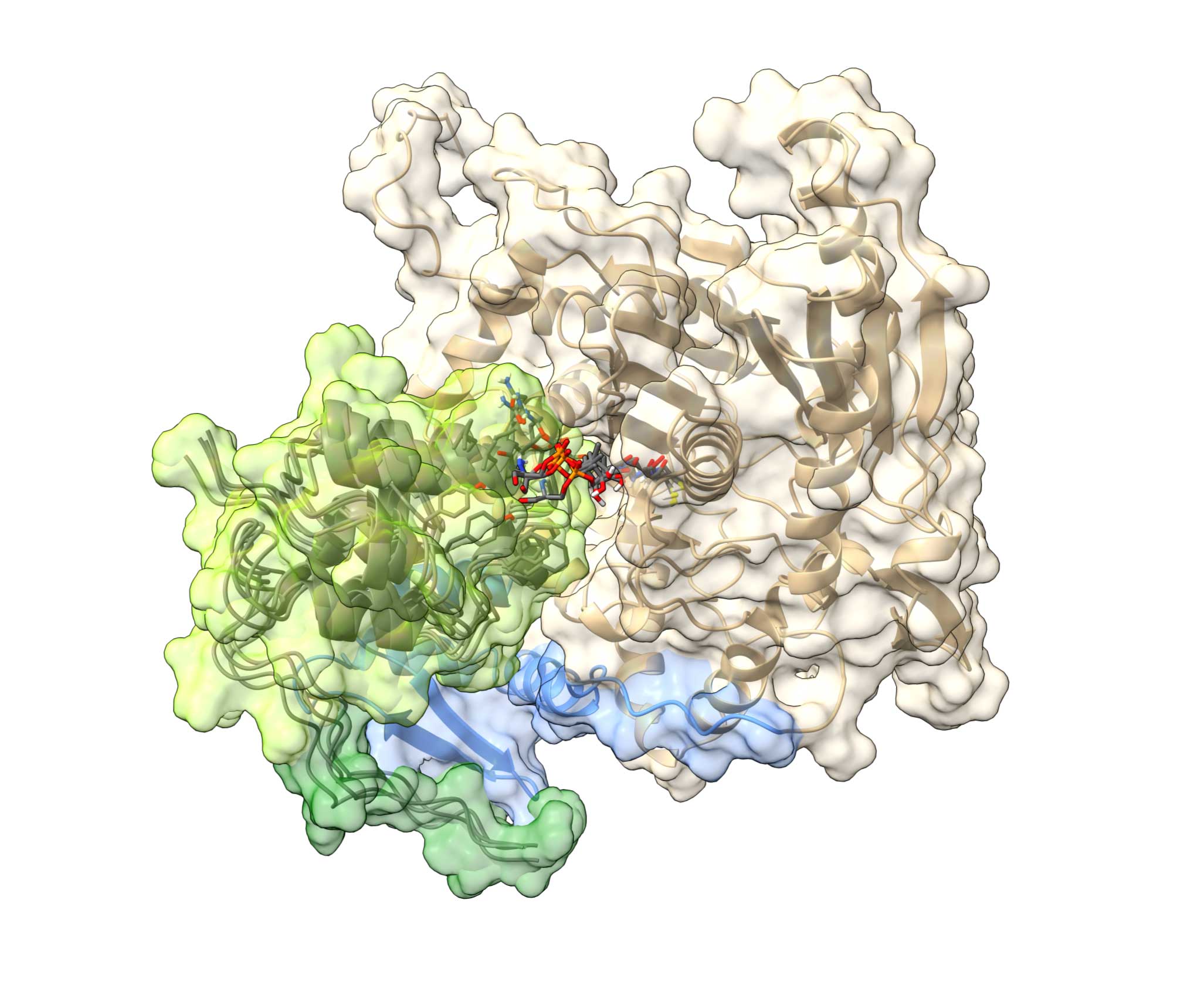 |
Structural biology and engineering of systems involved in natural products biosynthesisWe employ NMR-led integrated structural biology approach to study mechanistic details of various PKS/NRPS biosythetic pathways. Highlights:
|
NMR facilities
(NMR hall in Millburn House)
The group uses mostly 2 x 500 MHz (various MAS probes from 4 mm to 1.3 mm), 600 MHz (various MAS probes from 4 mm to 1.3 mm, a unique 0.8 mm 100 kHz probe from Ago Samoson; upgraded to a new Avance Neo console in 2021 funded by BBSRC ALERT19 BB/V50967X/1 ). We are also frequent users of the the UK High-Field Solid-State NMR Facility (850 MHz, 1 GHz and soon 1.2 GHz spectrometers).
(NMR hall in MAS building - left to right: solid-state NMR: 700 MHz WB spectrometer with 0.7, 1.3 and 3.2 mm MAS probes, solution NMR: 500 MHz, 600 MHz, 700 MHz with cryoprobe - upgrade to new console and TCI and TXO cryoprobes in 2023 funded by BBSRC ALERT21 BB/W020297/1).
The group is the main user of the ERC & UW funded 700 MHz solid-state NMR spectrometer and frequent users of 700 MHz and 600 MHz solution NMR spectrometers. More details on the facilities in the Department of Chemistry.