Deciphering the Molecular Mechanisms of Mammalian RNA Virus Replication
Principal Supervisor: Dr Jeremy Keown
Secondary Supervisor(s): Dr Nicole Robb
University of Registration: University of Warwick
BBSRC Research Themes: Understanding the Rules of Life (Microbiology, Structural Biology)
Apply now!
Deadline: 23 May, 2024
Project Outline
Bunyaviruses are an order of viruses containing over 550 unique members that infect humans, animals, and plants. Some bunyaviruses have been labelled by the World Health Organisation as posing a great risk to public health due to the lack of effective countermeasures and capacity to spread extensively. Additionally, bunyaviruses cause extreme economic and societal damage through their infection of livestock, crop plants, and wild animals. While there is extensive diversity in these viruses, they all perform common processes including entering a cell, copying their genetic material, and making mRNA to produce viral proteins. My group investigates how RNA viruses copy their genetic material (replication) and produce viral mRNA (transcription) and investigate the role of host proteins in controlling these processes.
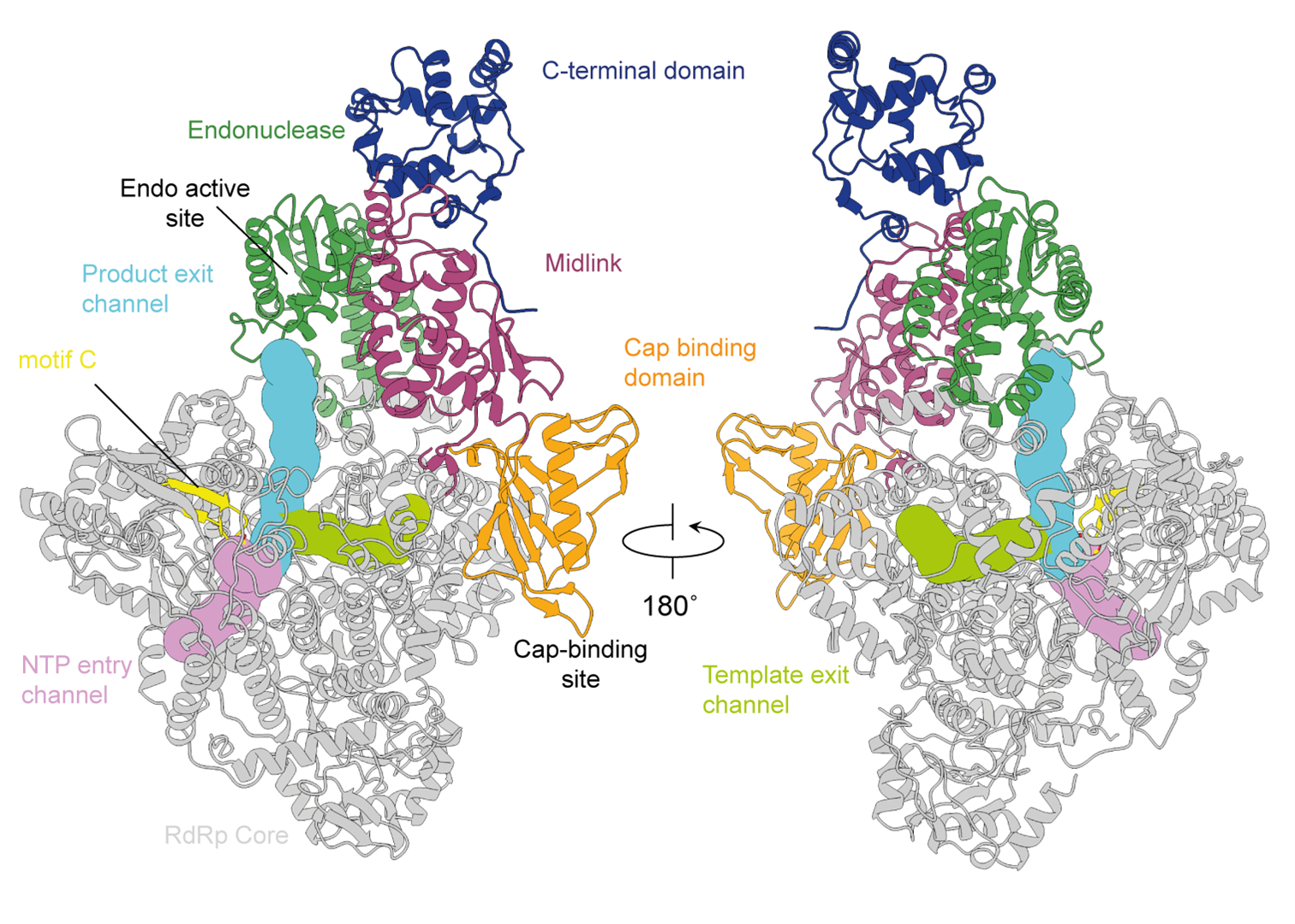
The genetic material, or genome, of a bunyavirus is formed by two or three segments of single stranded RNA, each RNA segment encodes the information to produce one viral protein. My group focuses on one of these proteins, the multifunctional viral RNA-dependent RNA polymerase. While all bunyaviruses contain a viral polymerase, the features of this polymerase differ between different viral families. Some viruses have a minimal undecorated polymerase, where others are almost double the size and contain additional domains that regulate polymerase function. Understanding how the polymerase functions is key to understanding how these viruses replicate. Projects in the lab focus on characterising bunyavirus polymerase using state-of-the-art structural biology techniques complimented with functional characterisation.
In the lab our current focus is on two families of viruses called the Nairoviridae and Hantaviridae. These families of viruses cause extensive harm to livestock and are now being discovered in new regions of the world as climate changes facilitates their spread. Transmission of viruses from animals to other animal species and humans is an increasing phenomenon. Stopping or reducing these outbreaks relies on a foundation of fundamental biological research. We believe by studying multiple virus families we can develop a broad and generalisable understanding of the bunyavirus biology and have begun to determine high resolution structures of Hantaviridae polymerase using cryoEM.
Using purified proteins and synthetic RNA we will determine structures of polymerase using cryoEM in conformations that mimic stages of transcription and replication. To characterise the function of the polymerase we will use both in vitro assays and minireplicon systems which recapitulate key stages of viral infection. As part of these projects, you will learn molecular biology, protein expression, single particle cryoEM, radionucleotide incorporation assays, and minireplicon functional assays. By combining these techniques, we will build a detailed picture of key stages in the viral lifecycle.
References
Keown, J.R., Carrique, L., Nilsson-Payant, B., Fodor, E., and Grimes, J. M. (2023) Structural Characterization of the Full-length Hantaan Virus Polymerase. BioRxiv. doi: https://doi.org/10.1101/2023.06.09.544421
Techniques
- Molecular biology and protein expression (E. coli and insect cells)
- Structural biology (X-ray crystallography and cryoEM)
- Mammalian cell culture
- Radionucleotide activity assays
- Mass spectrometry and proteomics