Using single molecule super-resolution microscopy to reveal how bacteria remodel their cell wall
Principal Supervisor: Dr Séamus Holden
Secondary Supervisor(s): Professor David Roper, Professor Phil Stansfield, or Dr Allister Crow depending on project undertaken
University of Registration: University of Warwick
BBSRC Research Themes: Understanding the Rules of Life (Microbiology)
Apply now!
Deadline: 23 May, 2024
Project Outline
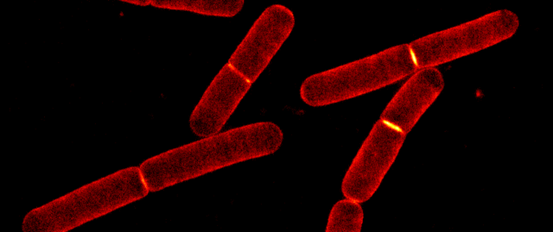
Bacteria are surrounded by a mesh-like cell wall which protects them from bursting due to their high internal pressure of turgor pressure. A better understanding of the fundamental principles of bacterial cell wall remodelling will help us to understand how bacteria resist antibiotics and may help identify new targets for antibiotic drug development.
In the Holden lab at Warwick University, we study the biology and biophysics of bacterial cell wall remodelling in Bacillus subtilis and Escherichia coli (see [1,2]). We currently focus on two closely related protein machineries: the elongasome, responsible for growing the bacterial cell wall by adding new material to it, and the divisome, responsible for dividing the cell by building a mid-cell crosswall (septum). We investigate how these nanoscale cell wall synthesis complexes precisely build and sculpt a comparatively massive micron-sized bacterial cell wall. This problem bridges microbiology, biophysics, and super-resolution microscopy.
Principal techniques used in our lab include:
- In vivo and in vitro single molecule fluorescence microscopy - to follow the activity of individual proteins directly in living cells as they build the cell wall or in vitro using reconstitution approaches
- Super-resolution microscopy - to image the organisation of proteins in vivo with nanoscale resolution
- Molecular microbiology, bacterial genetics and bacterial cell biology - to understand how the proteins work at the molecular level,
- Image analysis and data science - to analyse complex microscopy datasets.
About Our Lab
The Holden lab are a diverse group of biologists, biophysicists and microscopists who investigate fundamental mechanisms of bacterial cell envelope remodelling using advanced light microscopy (see e.g. [1,2]). We are based at the brand new £54M Interdisciplinary Biosciences Building at the School of Life Sciences, University of Warwick, which brings together researchers from microbiology, eukaryotic cell biology and neuroscience in a single interdisciplinary, collaborative environment with state-of-the-art facilities. Further information about the lab can be found at https://holdenlab.github.io/
We take a team science approach and are highly collaborative with a wide network of outstanding collaborators at Warwick and across the world including as far as Australia. During the PhD there will be substantial opportunity to engage in collaborative research.
Is This Project Right For You?
This project would suit students from either biology- or physics-allied disciplines, such as biochemistry, biomedical sciences or physics. The principal requirements for recruitment are outstanding potential, passion for fundamental science and a desire to work across disciplinary boundaries. Extensive interdisciplinary training will be provided and both the project and the training can be tailored to the initial skillset of the student.
What Will You Be Doing For Your PhD Research?
The final PhD project to fit broadly within the research directions described above and will be developed with the student according to their interests and skill set within a collaborative research environment. We urge students interested in the project to get in contact at earliest possible opportunity and engage in mini project opportunities ().
References
- K.D. Whitley, C. Jukes, N. Tregidgo, E. Karinou, P. Almada, Y. Cesbron, R. Henriques, C. Dekker, S. Holden, Nat. Commun. 12 (2021) 2448. https://doi.org/10.1038/s41467-021-22526-0.
- A.W. Bisson-Filho, Y.-P. Hsu, G.R. Squyres, E. Kuru, F. Wu, C. Jukes, Y. Sun, C. Dekker, S. Holden, M.S. VanNieuwenhze, Y.V. Brun, E.C. Garner, Science. 355 (2017) 739–743. https://doi.org/10.1126/science.aak9973
Techniques
- Single molecule fluorescence microscopy, super-resolution fluorescence microscopy (supervision training)
- Quantitative image and data analysis with Python and ImageJ (supervision training).
- Bacterial cell biology, bacterial genetics, molecular biology (supervision training)
- Collaborative projects could involve additional skills eg biochemistry (with Roper lab, Warwick), computational molecular dynamics (with Stansfeld lab, Warwick).