Circular Dichroism
Circular dichroism (CD) is the difference in absorbance of left and right circularly polarised light. For randomly oriented solutions CD is non-zero only for chiral molecules. CD can be used to probe the enantiomeric purity of small molecules (as long as the signal expected for 100% enantiomerically pure samples is known). More commonly CD is used to probe the structure of biomacromolecules such as proteins and DNA. CD is the ideal technique for estimating the secondary structure content of proteins. Our main applications of CD currently are to
(i) Determine the structure of proteins and nucleic acids as an adjunct to linear dichroism (LD) data collection, and
(ii) Analyse the structure of protein biopharmaceutical products. We operate a contract service for this purpose.
Our most recent CD research project has been to establish a new standard for calibrating CD spectropolarimeters that is chemically and enantiomerically stable, has bands at all wavelengths of interest and is available enantiomerically pure in both enantiomeric forms. This work will soon appear in Chirality.
Protein stucture analysis
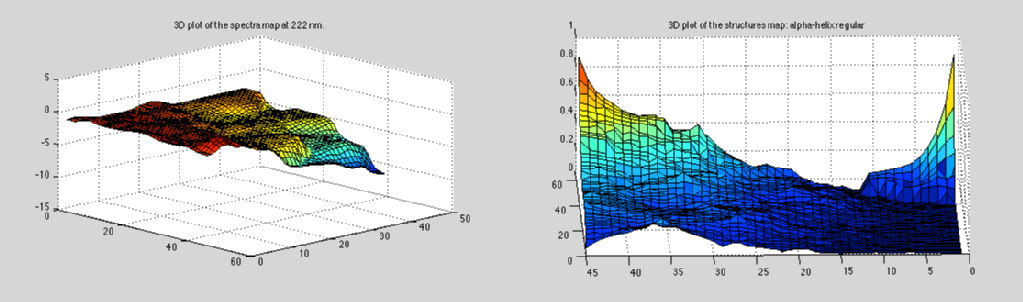
SSNN, or secondary structure neural network is a self organising feature map (SOFM) used to cluster data from circular dichroism to estimate protein secondary structures developed by Vincent Hall, Anthony Nash and Alison Rodger.
The program and the instructions are available from here.