Mini-Project 2
Cloning and Expression of Transcriptional Repressors in Escherichia coli
Supervised by: Dr. Christophe Corre
Following the sequencing of entire Streptomyces genomes, an unexpectedly large number of antibiotic-like gene clusters were found to be encoded. Many of these cryptic gene clusters (cryptic because their products are not known) were predicted to encode the biosynthesis of novel natural products. Under laboratory conditions, these cryptic biosynthetic gene clusters are often not expressed. The main reason is the antibiotic production is tightly controlled. The regulation of antibiotic production of several Streptomyces secondary metabolites is known to be triggered by low-molecular weight, diffusible, signalling molecules.
The γ-butyrolactone regulatory system triggers secondary metabolite production in the gram-positive, soil-dwelling, filamentous bacterial genus Streptomyces.The representative of the γ-butyrolactones is A-factor in S. grisues, which was the first to be found. A-factor induces the production of the antibiotic streptomycin. AfsA is an enzyme for A-factor biosynthesis and ArpA is the A-factor receptor protein that serves as a transcriptional factor by using A-factor as the ligand.
Recently a novel class of molecules that induce antibiotic production in actinomycete bacteria have been discovered called 2-alkyl-4-hydroxymethylfuran-3-carboxylic acids (AHFCAs). They are thought to interact with, and change the characteristics of, transcriptional repressor proteins that belong to the ArpA-subfamily of the TetR proteins. The TetR family of transcriptional repressors is well represented and widely distributed among bacteria. They have two domains:
• A DNA-binding domain that recognise specific DNA sequences in promoter regions
• A ligand binding domain that interact with specific molecules.
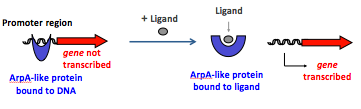
They strongly repress antibiotic and signalling molecule production. Upon binding of a signalling molecule, repressors are released for the promoter region of the target gene, which is then transcribed.
This project aimed at cloning genes that encode for 4 distinct ArpA-like proteins for two different sources Streptomyces avermitilis and Streptomyces venezuelae. These were be overexpressed in Escherichia coli and the solubility of the corresponding proteins will be investigated.A detailed understanding of the structure-activity relationships involved inArpA-like proteins with signalling molecules and DNA will provide new opportunities to manipulate antibiotic biosynthesis and potentially provide a means for discovering new compounds with clinical utility.
The spectrum below shows the FPLC trace of one of the transcriptional repressors S. venezuelae.
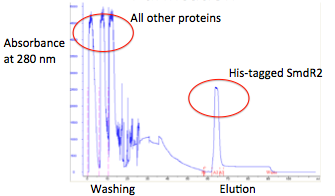
Transcriptional repressors can be effectively cloned and expressed in E. coli.
Use the links to view the poster and presentation produced for the assesement of this miniproject.
