Modelling noisy biochemical switches and networks
Supervisors: Timothy Saunders, Phillip Stansfeld
Biological systems often need to switch behaviour due to environmental (e.g. temperature) and other stimuli (e.g. cell-cell interactions). Yet, understanding how cells can make reliable decisions given the inevitable noisiness of the intracellular conditions remains an open problem. Here, we will develop stochastic models of small biochemical networks that undergo stochastic switching in behaviour. The project involves building a robust software framework for exploring the role of spatial and temporal fluctuations in protein number within different boundary constraints. Working closely with experimentalists (Loose lab, IST-Austria), we will generate predictions that can then be directly tested.
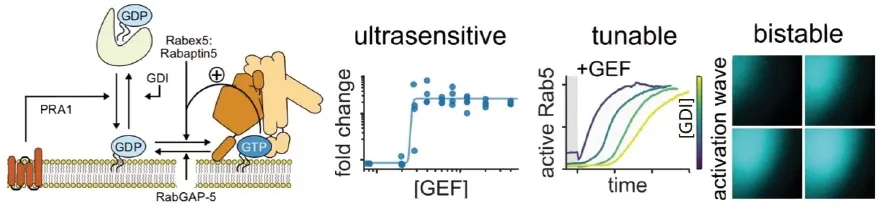
Biological systems are inherently noisy. Yet, they can perform complex operations robustly. How do cells make reliable biochemical decisions despite the inevitable presence of noise? Small GTPases are a large protein family that are critical in many cellular functions, such as growth and migration. They are activated at specific times and intracellular locations. Due to the high complexity of the living cell, the mechanisms that give rise to such high spatiotemporal control are difficult to study. An emerging theme is the importance of signalling cascades, where the presence of an activated GTPase leads to the recruitment and activation of another. We recently showed with in vitro reconstitution and stochastic simulations that a minimal GTPase network operates as an intrinsically bistable and stochastic switch (Bezeljak et al. PNAS 2020). This switch can be precisely tuned by different system inputs. Positive feedback in the network can even drive spatial patterns of activity.
In this project, we aim to use computational modelling to dissect how rapid and robust protein switches can operate effectively. In particular, we want to explore the interplay between stochasticity and the role of spatial constraints in governing behaviour in biochemical networks. This will entail developing novel spatial stochastic simulations to explore the role of geometry in affecting switching behaviour in small biochemical networks across multiple spatial and temporal scales. Creating efficient code that enables exploration of different network and spatial topologies is central to the project success. The modelling will be used to generate predictions – e.g. altering specific interactions – that can be tested experimentally by the Loose lab. Some time will be spent at IST-Austria, working alongside experimentalists.
Uncovering how spatial constraints affect protein network behaviour is an important step in understanding how different subcompartments of the cell respond differently to the same (noisy) biochemical inputs. Tackling such a problem is inherently multiscale, from single molecules to the cellular level. This interdisciplinary project offers an exciting opportunity for a student interested in the interface between stochastic simulations and biological application.
- Figure attached below. Legend: (Left) Outline of network interactions, (Middle) Ultrasensitive and tunable response of the network to different stimuli, (Right) Generation of waves of activation.