Glossary of terms
A
- AFLP
Amplified fragment length polymorphism.
A technique used to compare genomes and to identify and locate differences between them.
- Allele
Allele is the term used to describe one of two or more forms of a gene.
The word is derived from the shortening of the word Allelomorph.
Different alleles or allelic variants arise from mutation. They are usually found at the same position or place (locus) on homologous chromosomes. Two forms of a gene may be termed 'diallelism', and multiple forms 'multiple allelism'.
Different allelic forms may exist in a population, the frequency of their occurrence is measured by the 'allele frequency'.
Different allelic forms can have different effects on specific phenotypes
C
- CAPs marker
Cleavage Amplification Polymorphism.
A technique used to compare genomes and to identify and locate differences between them.
- Crop Domestication
Crop domestication and subsequent breeding efforts has resulted in modern crops with a restricted genetic base Sometimes referred to as ‘allelic canalisation’
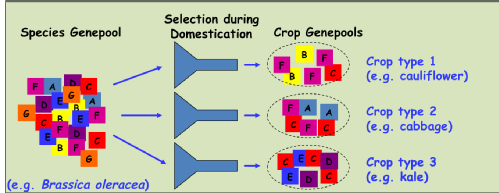
The selection process can lead to great variation in morphology between crop types but with overall reduced genetic diversity.
D
- Diploid
When there are two copies of each chromosome within each cell
- Diversity collection
A diversity fixed foundation set (DFFS) represents an informative set of genetically fixed lines representing a structured sampling of diversity across a genepool.
- DNA
Deoxyribonucleicacid
The genetic instructions to create most living organisms.
- Doubled haploid
When a haploid genotype undergoes chromosome doubling to produce homozygous lines in one generation. Doubled haploid plants are particularly useful for trait analysis since they are homozygous at all loci with reduced heterogeneity within lines. These 'genetically fixed' lines allow for replication of individuals during trials on different sites and over years.
G
- Gel electrophoresis
Molecules (DNA, RNA or protein) move or migrate through an electric field (electromotive force, EMF), with their movement restricted by a gel matrix, for example agarose, agar. The gel matrix is almost like a sponge in texture, with many small holes that the DNA/ RNA negotiates during its migration towards the negative pole (cathode). Molecules are sorted based on their size and charge, with large molecules moving more slowly.
A stain (e.g. ethidium bromide) is usually used to visualise the migrating molecules.
- Genetic marker
Genetic markers are used to identify different features in DNA sequence that can be used to differentiate between individuals in a population, or to classify individuals between different varieties or cultivars within a species.
- Genome
Total chromosomal DNA of an organism
- Genotype
The genotype of a plant is a word used describes the genetic make - up of the plant. The context that it is used depends upon whether it is being used to describe the whole genome, the DNA sequence of individual genes or a collection of scores at different genetic markers.
H
- Haploid
When there is one copy of each chromosome within a cell
- Heterozygous
Describes the situation when different alleles exist at a locus
- Homozygous
Describes the situation when the same alleles exist at a locus
L
- Linkage map
When all the members of the population have been scored (genotyped) with a set of molecular markers, the data can be used to make a linkage map. The linkage map describes the linear order of markers within linkage groups.
- Locus
The location of a gene on a chromosome (plural = loci)
M
- Marker assisted selection (MAS)
Also called marker aided selection refers to the process of using a marker type (in our case usually DNA based) to pre-select plants that have a specified genotype at that position in the genome. The marker(s) may be linked to a QTL or a gene that influence a particular target trait; therefore it is desirable to select only the plants that have the pre-defined genotype. This method allows hundreds of seedlings to be screened, and only the plants that have the desired target genotype to be retained.
P
- Phenotype
The phenotype of a plant is a term used to describe observable characteristics, such as height, biomass, leaf shape and so on.
Q
- QTL
Quantitative trait loci or locus
Positions in the genome statistically associated with a particular trait, in combination, that contribute towards the observed variation for a particular phenotype associated with a trait. Usually several genes are involved in the process.
- QTN
Quantitative trait nucleotide
A smaller more defined part of a QTL.
R
- Recurrent parent
The adapted cultivar used in a cross
- RNA
Ribonucleicacid
A similar molecule to DNA but involved in the manufacture of proteins.
S
- SNP
-
Single nucleotide polymorphism (SNP, pronounced snip)
DNA sequence variation - occurs when there is difference between individual nucleotide base pairs (A, T, C, G) in the genome (or other shared sequences) between members of a species, or paired chromosomes in an individual, Figure 1.

Figure 1 DNA molecule 1 differs from DNA 2 at a single base - pair location (in this case a C/ T polymorphism). Image from Wikipedia (wikki/Single-nucleotide-polymorphism).
In the example above the nucleotide Cytosine (C) has been replaced by the nucleotide Thymine (T).
- SSR
Simple sequence repeat
A genetic marker type, also called a “microsatellite”. These are used to classify DNA (genotypes) at defined positions in the genome facilitating comparisons between genomes of different plants.
T
- Trait
The word trait is used to describe a particular characteristic of a plant. It is usually applied to a defined characteristic that we wish to quantify. For example, the rate of post harvest head - yellowing in broccoli heads harvested from a population.
The attributes that describe a particular trait are well documented in many cases, allowing comparisons to be made by different researchers across the world.
V
- VeGIN
Vegetable Genetic Improvement Network
