Kinetochore dynamics
Introduction: Cell division is a highly complex 3+1D process, requiring extensive reorganisation of the cellular DNA achieved with high precision. The active machinery of chromosome separation is microtubules, a self polymerising protein that has an intrinsic dynamics, specifically it undergoes periods of growth and periods of retraction (catastrophes). These episodes are stochastic and related to the aging of the microtubule lattice. Whether this intrinsic dynamics underlies the semi-periodic oscillatory behaviour observed during division is a key question we hope to answer.
Microtubules are organised from the spindle poles and capture sister chromatids at the kinetochore, Fig. 1, ultimately pulling them apart to opposite poles of the cell. However, prior to separation, the kinetochores are aligned along the dividing plane and are observed to undergo semi-periodic oscillations. A number of proteins are known to affect the presence of oscillations, their period and their amplitude. These proteins, and their effects, will enable the elucidation of the mechanism of oscillations.
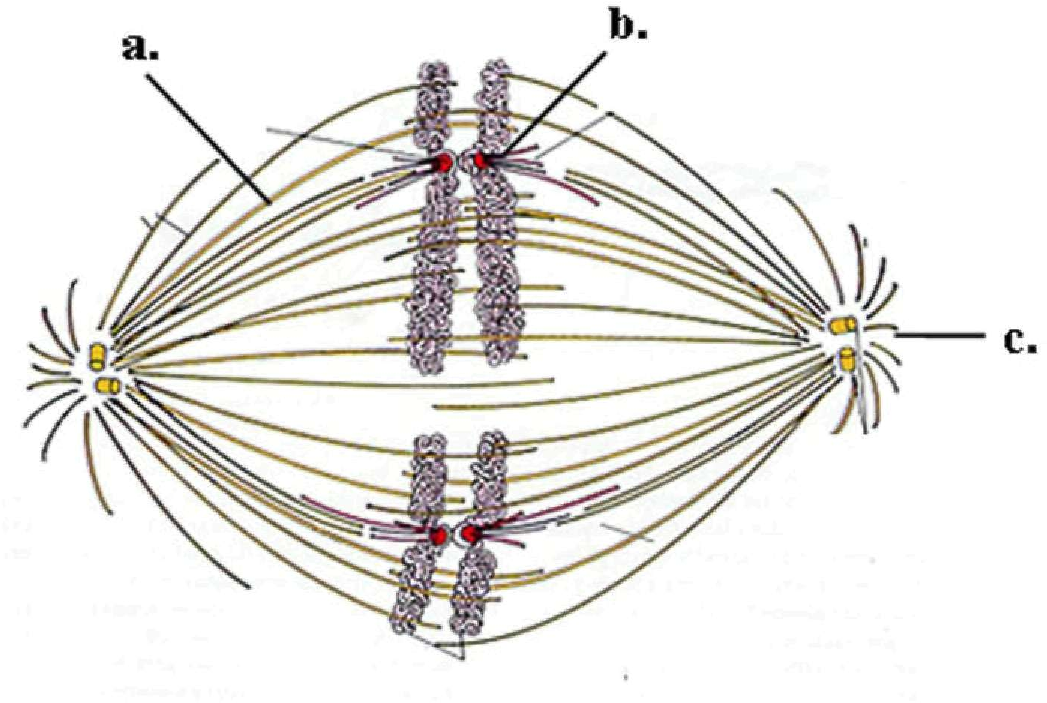 |
| Figure. a. microtubules, b. Kinetochore (red) attachment of microtubules to chromosomes, c. spindle poles. |
Data: Our experimental collaborators have refined their imaging techniques to obtain much higher spatial-temporal resolution of the kinetochore trajectories than could previously be achieved. They have also developed techniques to measure a number of parameters, eg tubulin turnover, and built a number of mutants. All data is on an human cell line. This data has shown that the dynamics is far more complex than previously thought requiring, and enabling, a far more detailing modelling and quantitative analysis programme.
Modelling: There are a number of models of the kinetochore structure and of the kinetochore-microtubule system that give rise to oscillations across different organisms, [1]. A primary aim of the project is to use these models to develop a model capable of explaining the new observations and determine the mechanism of oscillation. A number of approaches will be examined, firstly developing PDE versions of the existing models to enable dynamical systems techniques to be used. Secondly, we will use large scale discrete simulations.
Inference: A fundamental difficulty with parametrisation of these models is the complexity of the dynamics. A dedicated modelling avenue will also be developed alongside the modelling methods above, that utilise simpler mechanisms but capture the essential features of the dynamics. These models are built on Random walk processes (discretised sDEs) and thus can be inferred from the data using techniques such as Markov chain Monte Carlo methods. Since trajectories are short and thus insufficient to fully parametrise the more complex models, hierarchical modeling will be used to integrate across trajectories/data sets. Model selection techniques will be used to guide model development, ie ascertain which mechanisms are supported by the data.
Duties.
We will work with the experimental team lead by Andrew MacAinsh (Warwick Medical School) on the modelling and statistical analysis of kinetochore oscillations. We will be responsible for analysis of the experimental data, meeting target deadlines, and communicating results to the (experimental) team. This is a challenging but rewarding project tackling a major issue of key importance to the UK.
This project is funded through BBSRC

[1] Springs, clutches and motors: driving forward kinetochore mechanism by modelling. Elina Vladimirou, Ed Harry, Nigel Burroughs, Andrew D. McAinsh. Chromosome Research 2011 Apr;19(3):409-21. via PUBMED.
