Genetic markers: SNPs
The most common subunits of DNA are the four deoxyribonuclotides (nucleotides) containing the bases:
adenine (A), cytosine (C), guanine (G) and thymine (T).
A single Nucleotide Polymorphism (SNP) describes a single base difference between two
DNA sequences.
For example, a C/T substitution in the DNA of plant 2 compared to the same region of DNA in plant 1
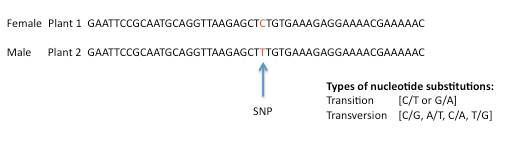
Using DNA sequence data from the parents of a population, SNPs can be identified
and used to screen their progeny – the individuals in the population.
For example 8 members of a population derived from a cross between two parents above.
|
C/T SNP scored in 8 lines
Plants 4 and 7 have inherited the polymorphism from the male donor. If the SNP was in the coding sequence of a gene this may alter the phenotype. |
Since the nucleotide is the smallest unit in DNA, SNPs represent the most abundant genetic marker type. New sequencing technologies enable thousands of SNPs to be discovered. These new markers have revolutionised the resolution attainable for genetic maps, and enhanced the precision of association analysis. SNPs used in genetic maps can be used to align sequence data to their respective chromosome positions. This is particularly useful in the absence of a fully assembled genome, and is essential during genome assembly and validation. |

