Metals in Medicine: Research
Organometallic anticancer complexes |
|||||
|
We are studying the design and mechanism of action of ruthenium and osmium arene anticancer complexes. Our research involves the synthesis and full characterization of new organometallic complexes, the study of their coordination chemistry and the determination of relevant kinetic and thermodynamic parameters. In addition, we study their reactivity towards biomolecules and determine their cytotoxicity both in vitro and in vivo. The principal techniques used to study these complexes are multidimensional, multinuclear NMR, (protein) X-ray crystallography, advanced mass spectrometry and cytotoxicity testing against various cancer cell lines. |
 |
||||
Ruthenium-Arene Complexes |
|||
|
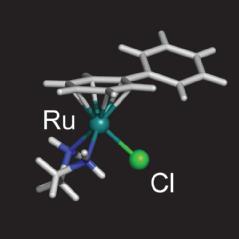
A typical half-sandwich ruthenium(II) anticancer complex |
|||
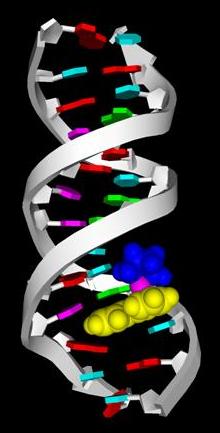
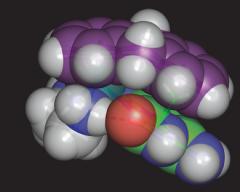 Multiple Interactions in 9-EtG binding to a Ruthenium-Arene |
|||
|
|
|||
|
|
|||
|
|
|||
Osmium-Arene ComplexesWe have recently extended this chemistry to osmium, the heavier congener of ruthenium. Completely different reaction kinetics are to be expected for the Os(II) arene complexes. The aqueous and hydrolytic chemistry of osmium analogues of the half-sandwich ruthenium(II) complexes were studied by systematic variation of the chelating ligand on osmium and was found to differ substantially from the analogous ruthenium complexes. The rational control of the chemical reactivity of the osmium complexes, however, did allow us to design cytotoxic osmium-arene complexes with promising activity against human A549 and A2780 ovarian cancer cells. |
|||
|
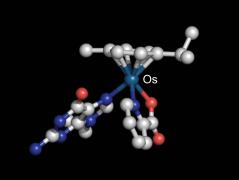
Nucleobase adduct of an osmium-arene half-sandwich complex |
| Selection of our recent publications on our organometallic anticancer complexes: | |||
1. |
Yan YK, Melchart M, Habtemariam A, Sadler PJ: Organometallic chemistry, biology and medicine: ruthenium arene anticancer complexes. Chemical Communications 2005:4764-4776 | ||
2. |
Morris RE, Aird RE, Murdoch PD, Chen HM, Cummings J, Hughes ND, Parsons S, Parkin A, Boyd G, Jodrell DI, et al.: Inhibition of cancer cell growth by ruthenium(II) arene complexes. Journal of Medicinal Chemistry 2001, 44:3616-3621. | ||
3. |
Habtemariam A, Melchart M, Fernandez R, Parsons S, Oswald IDH, Parkin A, Fabbiani FPA, Davidson JE, Dawson A, Aird RE, et al.: Structure-activity relationships for cytotoxic ruthenium(II) arene complexes containing N,N-, N,O-, and O,O-chelating ligands. Journal of Medicinal Chemistry 2006, 49:6858-6868. | ||
4. |
Wang FY, Habtemariam A, van der Geer EPL, Fernandez R, Melchart M, Deeth RJ, Aird R, Guichard S, Fabbiani FPA, Lozano-Casal P, et al.: Controlling ligand substitution reactions of organometallic complexes: Tuning cancer cell cytotoxicity. Proceedings of the National Academy of Sciences of the United States of America 2005, 102:18269-18274. | ||
5. |
Chen HM, Parkinson JA, Parsons S, Coxall RA, Gould RO, Sadler PJ: Organometallic ruthenium(II) diamine anticancer complexes: Arene-nucleobase stacking and stereospecific hydrogen-bonding in guanine adducts. Journal of the American Chemical Society 2002, 124:3064-3082. | ||
6. |
Chen HM, Parkinson JA, Morris RE, Sadler PJ: Highly selective binding of organometallic ruthenium ethylenediamine complexes to nucleic acids: Novel recognition mechanisms. Journal of the American Chemical Society 2003, 125:173-186. | ||
7. |
Liu HK, Berners-Price SJ, Wang FY, Parkinson JA, Xu JJ, Bella J, Sadler PJ: Diversity in guanine-selective DNA binding modes for an organometallic ruthenium arene complex. Angewandte Chemie-International Edition 2006, 45:8153-8156. | ||
8. |
Aird RE, Cummings J, Ritchie AA, Muir M, Morris RE, Chen H, Sadler PJ, Jodrell DI: In vitro and in vivo activity and cross resistance profiles of novel ruthenium (II) organometallic arene complexes in human ovarian cancer. British Journal of Cancer 2002, 86:1652-1657. | ||
9. |
Guichard SM, Else R, Reid E, Zeitlin B, Aird R, Muir M, Dodds M, Fiebig H, Sadler PJ, Jodrell DI: Anti-tumour activity in non-small cell lung cancer models and toxicity profiles for novel ruthenium(II) based organo-metallic compounds. Biochemical Pharmacology 2006, 71:408-415. | ||
10. |
Novakova O, Chen HM, Vrana O, Rodger A, Sadler PJ, Brabec V: DNA interactions of monofunctional organometallic ruthenium(II) antitumor complexes in cell-free media. Biochemistry 2003, 42:11544-11554. | ||
11. |
Novakova O, Kasparkova J, Bursova V, Hofr C, Vojtiskova M, Chen HM, Sadler PJ, Brabec V: Conformation of DNA modified by monofunctional Ru(II) arene complexes: Recognition by DNA binding proteins and repair. Relationship to cytotoxicity. Chemistry & Biology 2005, 12:121-129. | ||
12. |
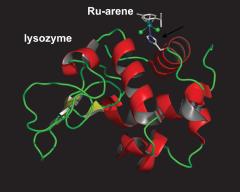
McNae IW, Fishburne K, Habtemariam A, Hunter TM, Melchart M, Wang FY, Walkinshaw MD, Sadler PJ: Half-sandwich arene ruthenium(II)-enzyme complex. Chemical Communications 2004:1786-1787. | ||
13. |
Yan YK, Melchart M, Habtemariam A, Peacock AFA, Sadler PJ: Catalysis of regioselective reduction of NAD(+) by ruthenium(II) arene complexes under biologically relevant conditions. Journal of Biological Inorganic Chemistry 2006, 11:483-488. | ||
14. |
Peacock AFA, Parsons S, Sadler PJ: Tuning the hydrolytic aqueous chemistry of osmium arene complexes with N,O-chelating ligands to achieve cancer cell cytotoxicity. Journal of the American Chemical Society 2007, 129:3348-3357. | ||