Nick R Waterfield
In the Waterfield group we are interested in understanding the molecular mechanisms and ultimately exploiting natural bacterial toxin systems for biotechnology purposes.
To do this, we routinely use cutting edge molecular biology and genomics techniques to rationally alter and edit natural systems such that they can perform tasks of interest to us. The Gram-negative pathogen Photorhabdus deploys a wide diversity of protein toxin delivery systems. We are currently studying the molecular mechanisms of several such systems. How are they targeted? Can they be adapted for medical or agricultural purposes? These projects are necessarily interdisciplinary and so we engage with inter-department projects. More specifically with Prof Matt Gibson; a polymer and biomaterials chemist and Prof Peter Scott; on novel antimicrobials, both from the Warwick Chemistry department.
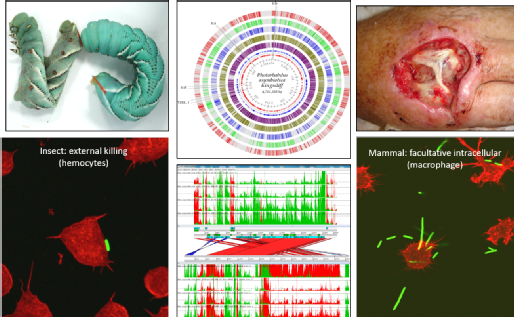
Another major focus in the group is Natural Product drug discovery, specifically from the insect pathogen Photorhabdus. Importantly, all natural products serve specific biological roles unlike the majority of molecules in synthetic compound libraries. If we can determine their activities we may exploit them for unmet medical or agrochemical needs. The majority of natural products produced by Photorhabdus are used to kill their insect host or combat competitors and predators, thus presenting a useful source of potential novel insecticides and antimicrobial drug candidates. In order to combat the rising threats of antibiotic and pest/parasite drug resistance, we need to develop novel compounds that exhibit new modes of action distinct from those of current drug targets. We are currently developing an imaginative trans-disciplinary methodological pipeline that will grant access to the wealth of cryptic natural products inferred from bioinformatic analysis of the genome sequences of the insect and human pathogenic genus Photorhabdus. These bacteria are sufficiently closely related to E. coli that many available synthetic biology genetic tools are applicable.
Selected publications:
1. Vlisidou, I., Hapeshi, A., Healey, J.R., Smart, K., Yang, G. and Waterfield, N.R., 2019. The Photorhabdus asymbiotica virulence cassettes deliver protein effectors directly into target eukaryotic cells. eLife, 8. doi: 10.7554/eLife.46259
2. Simpson, D.H., Hapeshi, A., Rogers, N.J., Brabec, V., Clarkson, G.J., Fox, D.J., Hrabina, O., Kay, G.L., King, A.K., Malina, J. and Millard, A.D., 2019. Metallohelices that kill Gram-negative pathogens using intracellular antimicrobial peptide pathways. Chemical Science. (selected as a 'Hot Article') doi: 10.1039/C9SC03532J
3. Chen, L., Song, N., Liu, B., Zhang, N., Alikhan, N.F., Zhou, Z., Zhou, Y., Zhou, S., Zheng, D., Chen, M. and Hapeshi, A., 2019. Genome-wide Identification and Characterization of a Superfamily of Bacterial Extracellular Contractile Injection Systems. Cell reports, 29(2), pp.511-521. https://doi.org/10.1016/j.celrep.2019.08.096
4. Hapeshi, A. and Waterfield, N.R., 2016. Photorhabdus asymbiotica as an insect and human pathogen. In The Molecular Biology of Photorhabdus Bacteria (pp. 159-177). Springer, Cham. https://doi.org/10.1007/82_2016_29
5. Yang, G. and Waterfield, N.R., 2013. The role of TcdB and TccC subunits in secretion of the Photorhabdus Tcd toxin complex. PLoS pathogens, 9(10), p.e1003644. https://doi.org/10.1371/journal.ppat.1003644
6. Yang G, Hernández-Rodríguez CS, Beeton ML, Wilkinson P, Waterfield NR. Pdl1 is a putative lipase that enhances Photorhabdus toxin complex secretion. PLoS pathogens. 2012 May 17;8(5):e1002692. https://doi.org/10.1371/journal.ppat.1002692
Co-workers
Joe Healey, Research Fellow
Max Addison, PhD Student
Duha Alkhde, , PhD Student
Amy Godfrey, PhD Student
Shathviga Manoharan, PhD Student
Grace Taylor-Joyce, PhD Student

