Protein Structure and Function
Protein structure research within WMS has several aims. These include:
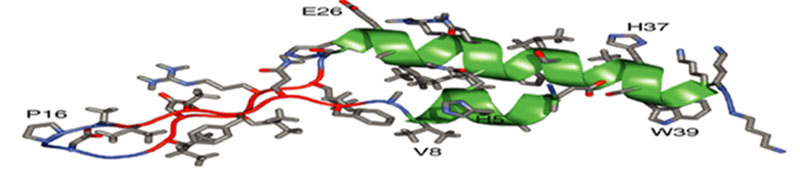
- Elucidating the detailed molecular structure of proteins by biophysical techniques, such as NMR spectroscopy, to further the basic understanding of the relationships between molecular shape and biological function
- Studying proteins the structure of which is not amenable to resolution by biophysical methods but which catalyse key metabolic processes, and are important targets in pharmacological strategies to prevent or alleviate disease
- Combining computational and experimental studies to study the mechanisms of folding and misfolding of membrane proteins and understanding, at the molecular level, how point mutations lead to protein misfolding
- Understanding the changes induced in protein structure through molecular damage induced by toxic metabolites in specific pathological states (e.g. diabetes, arthritis), to facilitate early diagnosis of these diseases through the development of novel biomarkers
- Studying the structural detail of protein – ligand interactions, to understand the networks of such interactions that exist naturally, the mechanisms underlying their malfunction in important metabolic diseases, and aid drug discovery. We perform combined computational and experimental studies to understand interactions of small molecules in numerous systems
We have strong links with:
- Major companies within the pharmaceutical and nutriceutical industries
- European consortia of universities and research centres
- Liverpool School of Tropical Medicine
- Global strategic alliances e.g. with the Universities of Monash (Australia) and Boston (USA)
- Departments within UoW: Systems Biology, Chemistry, Life Sciences, Engineering
We undertake research at the molecular end of translational research, with the aim of applying our observations to clinical scenarios, both in real time and in the long-term. We conduct research in the following areas:
- The structural and biophysical analysis of elements of the human immune innate system (e.g. the C-type lectins DC-SIGN and DC-SIGNR) that are expressed on dendritic cells and lymph node endothelial cells, and interact with glycosylated host adhesion molecules and viral glycoproteins.
- The mechanisms of folding and misfolding of the mammalian dim-light photoreceptor and G- protein coupled receptor, rhodopsin. Misfolding of rhodopsin is believed to be a primary cause in the most common inherited retinal degeneration disease, Retinitis pigmentosa, which ultimately causes blindness.
- The effects of protein damage by glycation, oxidation or nitration on the function and turnover of proteins in the plasma (e.g. the proteins associated with lipoproteins) and tissues (e.g. endothelial cells) of individuals with diabetes, arthritis and kidney disease. These studies enable the understanding of the accelerated deterioration of the function of tissues in chronic metabolic diseases.
- The intra-molecular and protein-membrane interactions of key proteins (carnitine palmitoyltransferases, diacylglycerol acyltransferases) involved in the metabolism of lipids in conditions such as hypertriglyceridaemia, hepatic steatosis and ketosis.
- The role of post-translational modification of proteins by phosphorylation, specifically that of the proteins KSR-1 (a scaffold protein that enables other proteins to interact with each other within specific cellular compartments), and of acetyl-CoA carboxylase 2, which regulates the utilisation of the major fuel (fatty acids) for key organs, including the heart.
- The refinement of a technique that enables the identification (using Surface Plasmon Resonance) of antibodies the pre-operative clearance of which enables the performance of renal transplantations using non-immune compatible donor organs.
- Coarse grained simulations of lipids in membranes, and their interactions with proteins, using the MARTINI force field e.g. to study the interactions of proteins with lipid droplets and carbon nanomaterials.
Our protein structure work is addressing the following issues:
- The effect of the overproduction of methylglyoxal in diabetic hyperglycaemia on the damage of lipoproteins, and the use of these changes as the basis for the development of novel biomarkers, using mass spectrometry. Studies are being performed on the effects of glycation, and of reactive oxygen and nitrogen species on the molecular structure of proteins (e.g. ion channels) that are important in the exacerbation of pain and cardiovascular disease in diabetes and kidney disease
- The molecular mechanisms underlying the changes in signal transduction in insulin resistance and cancer, through studies on proteins including KSR-1, dimentin, carnitine palmitoyltransfereases, diacylglycerol acytransferases and acetyl-CoA carboxylases. These studies are aimed at understanding the mechanisms underlying changes in fuel utilization by tissues that occur in diabetes, obesity and cancer, which are increasingly recognised as having overlapping causes
- The ability of the hyperglycaemia of diabetes to disturb the protein-protein interactions involving the glycobiome involved in the function of the innate immune system, resulting in the loss of immune function in diabetes
- The role of protein ligand interactions in the function of molecules such as the metabotropic glutamate receptors and binding of allosteric ligands chlorin and anthocyanins to rhodopsin. For HIV capsid, we have carried out an extensive ligand screen that has led to the identification of novel compound classes that modulate the oligomeric state of the capsid.
We receive funding from the following sources:
- Medical Research Council
- Biotechnology and Biological Sciences Research Council
- Technology Strategy Board
- British Heart Foundation
- EU Seventh Framework Programme for Research
- Collaborations with foreign national research organisations
