Biomolecular NMR
Why solid-state NMR for biomolecular systems?
Understanding structural dynamics, or evolution of molecular structure in time, is essential for understanding molecular-level mechanistic details of biological processes. Solid-state NMR is a powerful tool to probe structure and dynamics of biomolecules in a site-specific manner. Its advantage is that it can be applied to insoluble systems and that it does not require long-range order to yield atomic resolution information. This makes solid-state NMR highly complementary to solution NMR and x-ray crystallography, especially for systems like amyloid fibrils and membrane proteins. In addition, the NMR line widths in the solid state are not intrinsically dependend on the size of the studied systems, which means that solid-state NMR can be used to characterize biomolecules in even very large assemblies on the order of hundreds of kilodaltons to megadaltons. Finally, solid-state NMR provides a very rich view of molecular motions occuring on timescales spanning nine orders of magnitude.
What do we do?
Our group both develops methodology for studying structure and dynamics of biomolecules and applies solid-state NMR techniques (often combined with other biophysical methods) to a wide variety of biological systems.
Method development
The main themes for method development include tools for quantifying protein motions, approaches to characterising large biomolecular complexes and methodology involving fast magic angle spinning.
Applications
The systems studied include large protein complexes (e.g. protein-antibody complexes, protein complexes from polyketide synthases, complexes involving membrane proteins), fibrils and membrane proteins in Lewandowski group and isotopically labelled intact plants in Dupree and Brown groups (example publication).
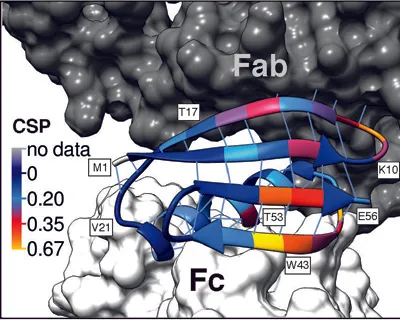 |
15N Chemical Shift Perturbations (CSP) for GB1 in a precipitated complex with IgG and GB1 free in solution projected onto the structure of GB1 in a model of the complex. Data from Lamley et al. JACS 2014. |
