Second level SnPM analysis for FSL users
|
|
Second level SnPM analysis for FSL users |
This page... introduction example data background design setup computation viewing results SnPM pages... manual PET example fMRI example FSL users |
OverviewThe purpose of this section is to supply the steps necessary to carry out a second level SnPM analysis using the results from an analysis carried out using the FMRIB Software Library (FSL). Specifically, instructions are given that will show the user how to select the contrasts of parameter estimates (copes) from FSL using SnPM. The individual copes can either be obtained from the individual analyses or from a group analyses that used all individuals of interest.If only the single subject analyses have been performed, the copes from all subjects must be registered to the same atlas space before reading into SnPM. If a group analysis has already been performed, then the individual copes have been registered to the same atlas space. FSL 3.0 and 3.1, SPM2, and SPM99 read Analyze format. By default SPM2, though, only allows a single volume per Analyze file, while FSL routinely uses multiple volume files. This becomes a problem only if a SnPM analysis is applied to a 2nd level FEAT directory, as the multiple subject's copes are all stored in a single multivolume image file. If the copes are coming from separate single-subject first level FEAT analyses, then there is no need to read multivolume Analyze files. The details of how to deal with multivolume files are discussed below. Note that FSL3.2beta creates NIFTI files, as does SPM5. Currently SnPM has no support for NIFTI files. |
Using Single-Subject FEAT directories
If you would like to run your SnPM analysis using the original single-subject FEAT directories, you first need to create copies of the copes and varcopes that are in the standard atlas space. This can be done using the featregapply on each single-subject FEAT directory.
featregapply <feat-directory-name> creates a reg_standard directory in the feat directory. In reg_standard is a stat subdirectory that contains copies of the copes and varcopes in the standard atlas space, one for each contrast.
Reading in the data
Once you've run featregapply on all .feat directories of interest you are ready to run SnPM! You can follow the steps for an fMRI analysis and when selecting cope images go to the *.feat/reg_standard/stats directory for each FEAT analysis and you will find the transformed copes and varcopes, as shown below.
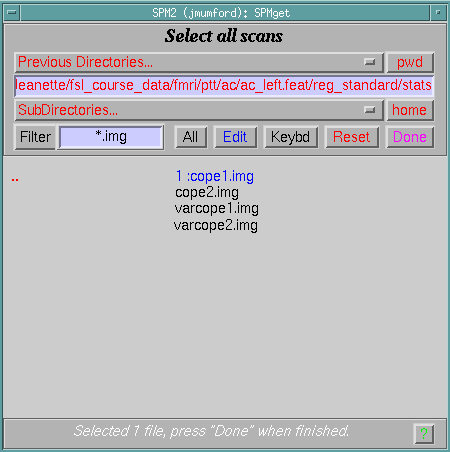
Using Group FEAT directories
After a group FEAT analysis is run a .gfeat directory is created which, among other things, includes a copy of the first-level copes for each subject in the form of a multivolume analyze volume.Each cope is located directly in the .gfeat directory and is labeled as cope#.img, where # is the number of the cope. The default settings of SPM are not set to recognize multiple volumes and hence if the cope#.img file is read into SPM without changing the defaults it will mistake the .img file as one subject instead of multiple subjects. The SPM defaults can either be changed for a single session or permanantly so that multivolume analyze volumes are recognized.
Changing SPM settings to allow for multivolume analyze volumes
Change for single session
To change the defaults for a single session, must select 'defaults' found on the bottom of the main SPM2 gui.You will then be asked which defaults area you are interested in and you want to select 'Miscellaneous Defaults'. Select the first two choices ('Log to file?' and 'Command Line input?') however you like. The third choice is 'Allow multi-volume Analyze files?' and for this you should select 'yes'. There is one more item to select after this and choose whatever value you like.
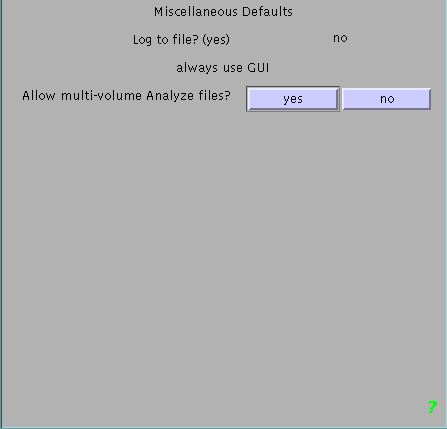
Note, this will only work for the current session of SPM. If you close and reopen SPM, you will need to reset the default to read multi-volume analyze files.
Change Permanently
If you would rather change the SPM defaults so that it will always recognize multi-volume analyze files, you will need to alter the spm_defaults.m file directly. Within this file, find the 'File format specific' section (shown below) and change defaults.analyze.multivol=0 to defaults.analyze.multivol=1.% File format specific %======================================================================= defaults.analyze.multivol = 0;
Reading in the data
SPM will now recognize multiple volume data and you can follow the steps for a fMRI analysis. When you want to select the images, look in the .gfeat directory and you will see that each cope has mutiple files, similar to what is shown below. If your cope#.img file is not follwed by comma and a number, first check that you did step 2 correctly and then check that your fsl analysis included all of the subjects you wanted.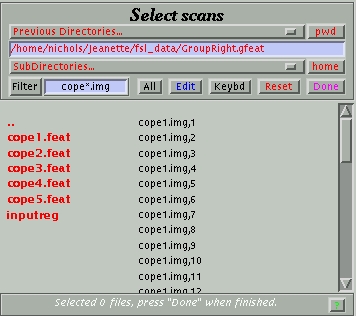
You may get an error message similar to the following, which may be ignored.
Warning: Assuming a scalefactor of 1 for "*/GroupRight.gfeat/cope1.img".
Back to main SnPM page
SnPM by Andrew Holmes of Astra Zeneca
& Tom Nichols of University of Michigan Biostatistics department
- SnPMauthors at <snpm-authors at umich.edu>
Neuroimaging
Statistics
Contact Info
Room D0.03
Deptment of Statistics
University of Warwick
Coventry
CV4 7AL
United Kingdom
Tel: +44(0)24 761 51086
Email: t.e.nichols 'at' warwick.ac.uk
Web: http://nisox.org
Blog: NISOx blog
Handbook of fMRI Data Analysis by Russ Poldrack, Thomas Nichols and Jeanette Mumford


![[Book Cover]](https://warwick.ac.uk/fac/sci/statistics/staff/academic-research/nichols/publications/books/handbookfmri_cover.jpg?maxWidth=125)