Epidemic outbreaks caused by environment, not evolution
5 August 2014
 Researchers have traced genetic changes in a bacterial pathogen over 450 years, and claim that epidemics of bacterial disease in human history may be caused by chance environmental changes rather than genetic mutations.
Researchers have traced genetic changes in a bacterial pathogen over 450 years, and claim that epidemics of bacterial disease in human history may be caused by chance environmental changes rather than genetic mutations.
In a study published in PNAS, a team led by the University of Warwick analysed 149 genomes of Salmonella enterica serovar Paratyphi A, which is a major cause of enteric fever. Enteric fever is currently estimated at 27 million clinical cases each year, resulting in 200,000 deaths.
Lead author, Zhemin Zhou from Warwick Medical School, said: “When epidemics break out, many scientists suspect they have been driven by increased virulence or fitness, possibly associated with the gain of novel genes or mutations. We wanted to trace genetic changes in a prominent bacterial pathogen back to see if this was true.”
The team reconstructed the genealogy, global transmission history and evolutionary history and found the pathogen originated at least 450 years ago, and had not changed dramatically over the centuries. This suggests that the pathogen had not become more efficient at causing enteric fever.
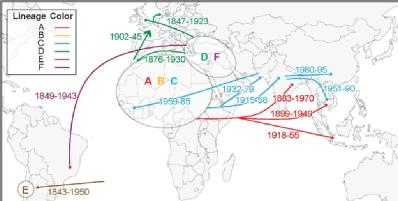
Zhemin Zhou said: “We found the pathogen formed seven distinct lineages that spread globally since the mid-19th century. Tracing the pathogen, we found there were genetic mutations that may have transiently improved drug resistance or improved the metabolic efficiency. However, most mutations were short-lived and removed by evolutionary forces.”
Senior author Professor Mark Achtman, Warwick Medical School, said: “We interpret the history of Paratyphi A as reflecting drift rather than progressive evolution. Our results indicate the crucial genomic contents of Paratyphi A, that cause enteric fever in humans, accumulated very early in its history.
“This implies that many epidemics and pandemics of bacterial disease in human history reflected chance environmental events, including geographical spread and/or transmission to naïve hosts, rather than the recent evolution of particularly virulent organisms.”
The study has been completed in collaboration with colleagues at University College Cork, Institut Pasteur, Paris, Public Health England and the Wellcome Trust Genome Campus in Cambridge.
Notes to editors:
- Image (pictured): The seven lineages of Paratyphi A and associated geographic transmissions with date periods.
- The paper, ‘Transient Darwinian selection in Salmonella enterica serovar Paratyphi A during 450 years of global spread of enteric fever’, is published in PNAS
Contact
Kelly Parkes-Harrison: Senior Press and Communications Manager, University of Warwick
Email: k dot e dot parkes at warwick dot ac dot uk
Phone: 02476 150868
Mobile: 07824 540863
