Andrew Bowman
Summary
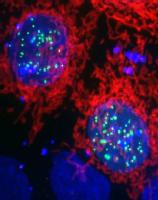
My main interest is focused on how eukaryotic DNA is packaged and organised within the cell. In particular, I am interested in how the core histones are assembled into nucleosomes to create a platform for higher order chromatin structure. As many nuclear processes require access to the genome to exert their function, such as transcription, DNA replication & DNA repair, correct assembly and disassembly of chromatin structure is imperative for proper cellular function.
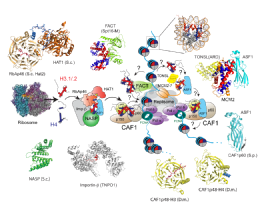
Key to the chromatin assembly process is a set of histone specific chaperoning proteins that govern histone folding, storage and incorporation into chromatin. These are very dynamic processes, with histones being incorporated into chromatin within minutes of being synthesised in the cytoplasm. During this time it is thought that histones are transferred between a number of different chaperoning and histone modifying proteins in a maturation process that produces deposition-competent histones.
I am interested in observing such transitions in living cells using the core histone H3 and H4 as models for chromatin assembly. In order to visualise and quantify such dynamic behaviour in living cells I am developing methods that interface synthetic biology and protein engineering tools with state of the art live cell imaging approaches.
It is hoped that being able to visualise these processes in real-time will broaden our understanding of one of the most fundament processes in chromatin biology.
Selected publications:
Mamar, H., Fajka-Boja, R., Mórocz, M., Jurado, E.P., Zentout, S., Mihuţ, A., Kopasz, A.G., Mérey, M., Smith, R., Sharma, A.B., Lakin, N.D., Bowman, A.J., Haracska, L., Huet, S., Timinszky, G., (2024). The loss of DNA polymerase epsilon accessory subunits POLE3-POLE4 leads to BRCA1-independent PARP inhibitor sensitivity. Nucleic Acids Res. https://doi.org/10.1093/nar/gkae439
Pardal A.J. & Bowman A.J. (2022). A specific role for Importin-5 and NASP in the import and nuclear hand-off of monomeric H3. eLife. e81755. doi: 10.7554/eLife.81755Link opens in a new window
Pardal A.J., Fernandes-Duarte F., Bowman A.J. (2019). The histone chaperoning pathway: from ribosome to nucleosome. Essays Biochem. 63(1):29-43.Link opens in a new window Review
Apta-Smith M.J., Hernandez-Fernaud J.R., Bowman A.J. (2018). Evidence for the nuclear import of histones H3.1 and H4 as monomers. EMBO J. doi:10.15252/embj.201798714Link opens in a new window
Bowman A. et al., (2017). "sNASP and ASF1A function through both competitive and compatible modes of histone binding." Nucleic Acids Res. (2017) 45(2):643-656Link opens in a new window
Bowman A, Lercher L, Singh HR, Zinne D, Timinszky G, Carlomagno T3 Ladurner AG. (2016). "The histone chaperone sNASP binds a conserved peptide motif within the globular core of histone H3 through its TPR repeats." Nucleic Acids Res 44(7):3105-17.Link opens in a new window
Bowman, A., C. M. Hammond, A. Stirling, R. Ward, W. Shang, H. El-Mkami, D. A. Robinson, D. I. Svergun, D. G. Norman and T. Owen-Hughes (2014). "The histone chaperones Vps75 and Nap1 form ring-like, tetrameric structures in solution." Nucleic Acids Res 42(9): 6038-6051.Link opens in a new window
Hondele, M., T. Stuwe, M. Hassler, F. Halbach, A. Bowman, E. T. Zhang, B. Nijmeijer, C. Kotthoff, V. Rybin, S. Amlacher, E. Hurt and A. G. Ladurner (2013). "Structural basis of histone H2A-H2B recognition by the essential chaperone FACT." Nature 499(7456): 111-114Link opens in a new window.
Bowman, A. and T. Owen-Hughes (2012). "Sulfyhydryl-reactive site-directed cross-linking as a method for probing the tetrameric structure of histones H3 and H4." Methods Mol Biol 833: 373-387.Link opens in a new window
Bowman, A., R. Ward, N. Wiechens, V. Singh, H. El-Mkami, D. G. Norman and T. Owen-Hughes (2011). "The histone chaperones Nap1 and Vps75 bind histones H3 and H4 in a tetrameric conformation." Mol Cell 41(4): 398-408.Link opens in a new window
Bowman, A., R. Ward, H. El-Mkami, T. Owen-Hughes and D. G. Norman (2010). "Probing the (H3-H4)2 histone tetramer structure using pulsed EPR spectroscopy combined with site-directed spin labelling." Nucleic Acids Res 38(2): 695-707.Link opens in a new window