Amira Ksibe
|
|
 |
Background
I have completed my Master degree in the chemistry department at Warwick University, 2011, Link  to Amira's AS:MIT wepage, studing zinc-binding ligands. Then in 2012, I have started my PhD research project entitled "Zinc on the move: Insights towards understanding zinc homeostasis in the open ocean cyanobacterium Synechococcus sp. WH8102".
to Amira's AS:MIT wepage, studing zinc-binding ligands. Then in 2012, I have started my PhD research project entitled "Zinc on the move: Insights towards understanding zinc homeostasis in the open ocean cyanobacterium Synechococcus sp. WH8102".
Research interests
In an attempt to resolve how open ocean cyanobacteria persist in regions where the zinc concentration is thought to be limited, I have hypothesised that analogy to siderophore-mediated iron acquisition and chalkophore-mediated copper acquisition, that the cyanobacterium Synechococcus sp. WH8102 might provide an extracellular zinc scavenger for the acquisition of this essential element. Therefore, I have developed an experimental method to isolate and purify the putative biogenic zinc-binding ligand (zincophore), using SPE cartridges composed of polystyrene-divinylbenzene resin and liquid chromatography. Then, I performed zinc limitation and repletion experiments on axenic cultures, and determined that this strain mounted an adaptive response for zinc under deplete and replete conditions, resulting in the induction and/or repression of a number of proteins.
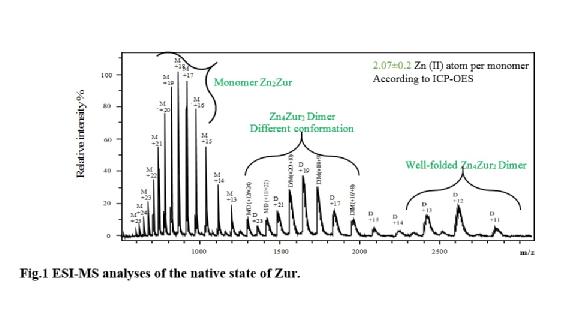
In the second part of PhD, I have sub-cloned and purified Zur, a zinc uptake regulator, synw_2401 gene, from Synechococcus sp.WH8102 in the absence of zinc ions in the cyanobactrium cultures. Then I have used mass spectrometry to probe the stoichiometry of zinc binding and the oligomeric state of this protein (Fig.1). Mass spectrometry confirmed that Zur existed as both a monomer with two zinc ions, Zn2Zur, and a dimer/well-folded dimer containing four bound zinc ions, Zn4Zur2 and this was in agreement with ICP-OES (Fig.1).
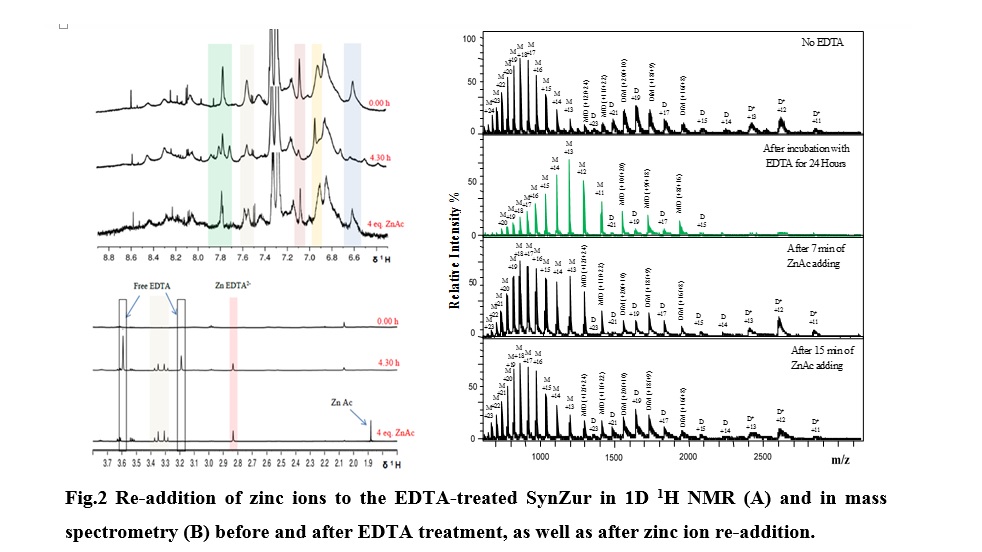
Treating Zur protein with EDTA resulted in an efficient removal of zinc ions from one site, the regulatory site, giving 0.9 ± 0.3 Zn(II) atoms per monomer. Adding back zinc ions to the EDTA-treated Zn1Zur species under anaerobic conditions yielded a comparable spectrum in NMR and ESI-MS to the native one (Fig.2). The new spectrum also displayed monomer, dimer as well as a well-folded dimer form containing 1.9 ± 0.4 Zn(II) atoms per monomer (using ICP-OES measurements).
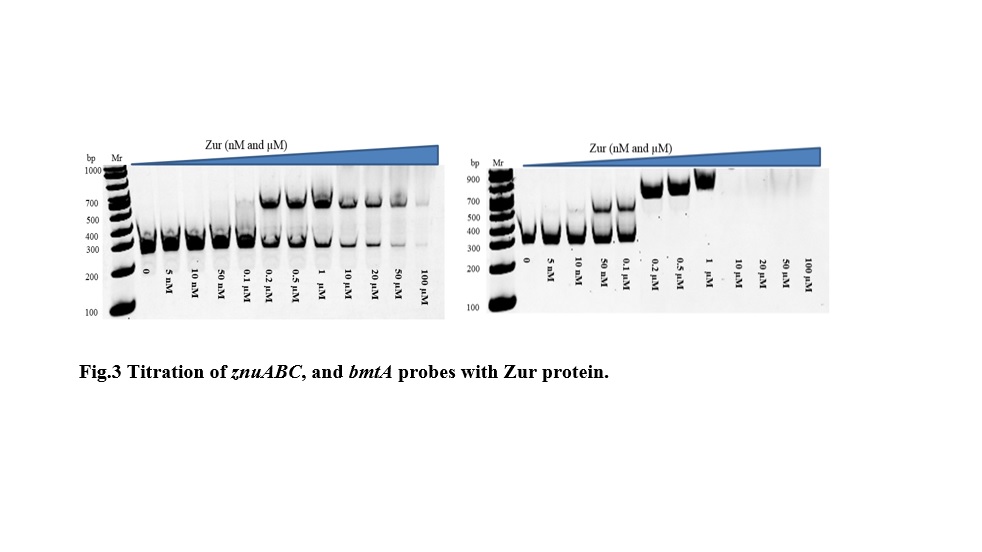
I have then used a gel electrophoretic mobility shift assays (EMSA) to test the binding of the Zur protein to predicted promoters, including those for znuABC system, bmtA (Fig.3).
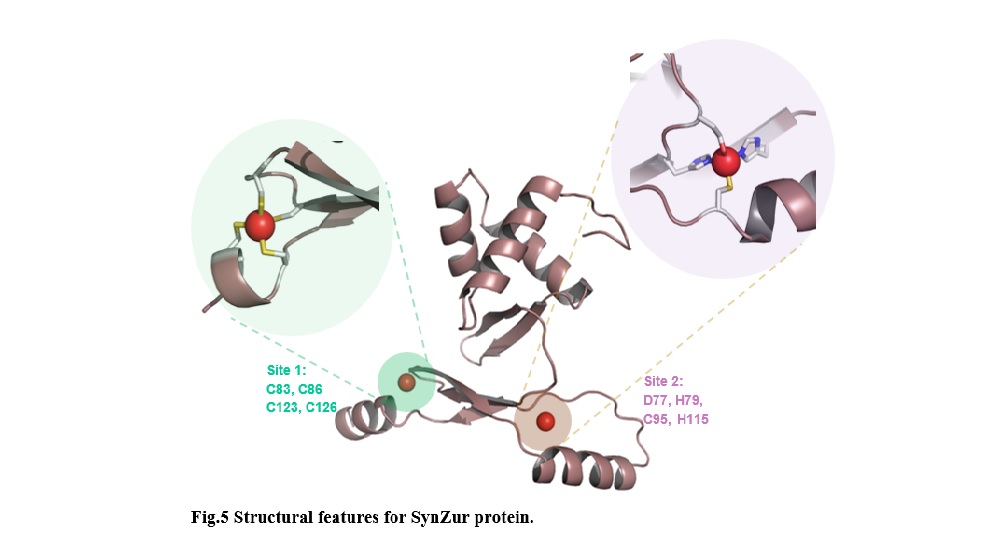
Waiting the structural determination of Zur protein in x-ray crystallography (Fig.4), I have created a 3D
homology modelling (Fig.5).