Pneumonia virus of mice
Pneumonia virus of mice (PVM)
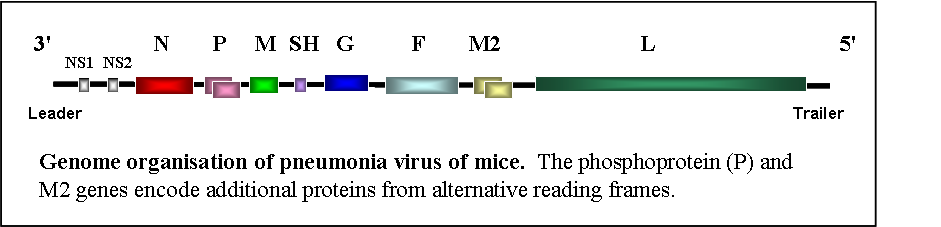
Members of the subfamily Pneumovirinae of the family Paramyxoviridae, given the collective name pneumoviruses, are responsible for acute respiratory infections in their hosts. The subfamily is divided into two genera on the basis of differences in genome organisation. The genus pneumovirus contains two members, the respiratory syncytial viruses, of which there are strains which infect humans, cattle (bovine RSV) and goats (caprine RSV), and pneumonia virus of mice (PVM) a common pathogen in laboratory animal colonies which infects a wide range of rodents and other animals, as well as man. PVM is the nearest phylogenetically related virus to human respiratory syncytial virus with which it shares a common genome organisation.
We, together with colleagues at NIH and Utrecht, have characterised several aspects of PVM including studies on the basic molecular biology and immunological characteristics of the disease in mice. Over recent years the technique of reverse genetics for negative strand RNA viruses has permitted the analysis of the function of virus proteins and their role in pathogenicity. Reverse genetics of negative sense RNA viruses permits the precise mutation of a cDNA copy of the virus genome which is subsequently rescued into infectious virus to determine the biological consequence of the mutation. In this way it is possible to study the function of specific proteins during the replication cycle and their influence on pathogenicity. Recently, reverse genetics systems have been developed by us and others. We are using this system to further investigate the molecular biology and pathogenesis of PVM.
