Bacterial canker of cherry
The disease
Bacterial canker is one of the most important diseases of cherry (Prunus avium L.). It is a major limitation for timber production from wild cherry and can limit the production of sweet cherry. It can also attack other Prunus species including plum (Prunus domestica).
Symptoms include cankers on twigs, branches and/or trunk, gum exudation, dieback and leaf spots.

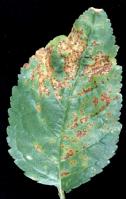
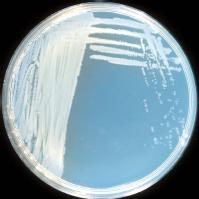
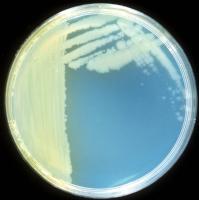
This disease can be caused by two pathovars of Pseudomonas syringae: pv. morsprunorum (Psm) and pv. syringae (Pss). Initial studies on sweet cherry attributed the disease mainly to Psm in the UK and to Psm and Pss in other countries; more recently Pss and/or intermediate forms between Psm and Pss were also found in sweet and wild cherry in the UK. We have carried out a survey of woodland plantations and nurseries in 2000/01 (Vicente et al. 2004. Eu J Plant Pathol. 110: 337-351).
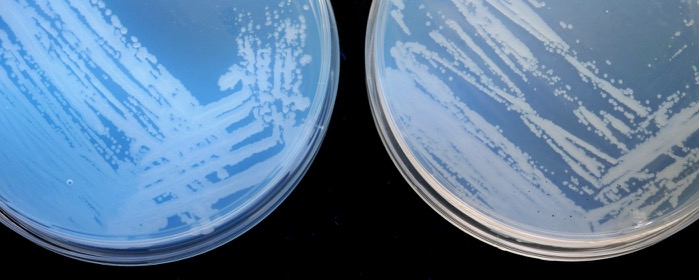
Discrimination of P. syringae isolates from cherry
The studies carried out at HRI were funded by Defra.
A collection of isolates from sweet and wild cherry was characterised by
- physiological and biochemical tests
- serological tests including agglutination and indirect-ELISA with polyclonal antisera
- pathogenicity tests using micropropagated plantlets of cherry and lilac
- rep-PCR fingerprinting
Physiological and biochemical tests can discriminate Psm races 1 and 2 from other P. syringae isolates. Agglutination and indirect-ELISA showed that Psm race 1 and race 2 are very uniform, but indicated a high variability amongst other P. syringae isolates (Vicente et al. 2004. Eu J Plant Pathol. 110: 337-351). The results of rep-PCR also showed that the Pss isolates are highly variable and the Psm isolates are generally very uniform within each race (Vicente and Roberts. 2007. Eu J Plant Pathol. 117: 383-392).
Pathogenicity tests are still necessary to discriminate the pathogenic Pss isolates.
Download handout of a poster and abstract presented at the 11th International Conference on Plant Pathogenic Bacteria, Edinburgh, Scotland 11-14 July 2006.
We have collaborated with NIAB EM (East Malling), the University of Reading and Imperial College, to compare whole genome sequences of representative strains of Pss, Psm and non-pathogenic Pseudomonas strains from cherry. This work has been published in Hulin et al. (2018) New Phytologist. doi: 10.1111/nph.15182
For more information, please contact Joana G. Vicente.
