Keith Leppard - Research in Detail
If you don't have a background in bioscience research, you may prefer to read an alternative summary with some simplified explanations.
From the moment they enter a host cell, viruses are modifying the environment to produce a favourable outcome for the virus: host responses that would slow or block infection are inhibited while host processes required for the virus life cycle are adapted and re-purposed. The overall focus of the laboratory is to understand these virus:host interactions and so to be able to manipulate them. We have focused particularly on human adenovirus type 5, a respiratory pathogen that causes only mild clinical symptoms and which has been extensively characterised in the past to provide a wealth of experimental tools and reagents.
PML proteins in virus infection and innate immunity
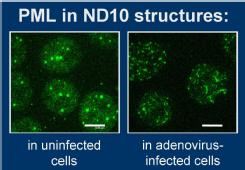
Of particular interest at present is the interaction between viral proteins and host nuclear structures known as PML nuclear bodies (PML-NBs). These structures are spherical multiprotein assemblies with a major component being the promyelocytic leukaemia protein, PML; this protein is produced by a gene that is the target of a reciprocal translocation in a large majority of cases of this leukaemia. PML-NBs are disrupted upon Ad5 infection, due to the action of the viral E4 Orf3 protein. We are interested to understand the molecular target for Orf3 within PML-NBs, the relevance of this disruption function to the outcome of infection and, through these studies, to learn about the role of PML-NBs within uninfected cells. There is evidence to link them, or their components, with a wide range of key cell processes, including innate immunity.
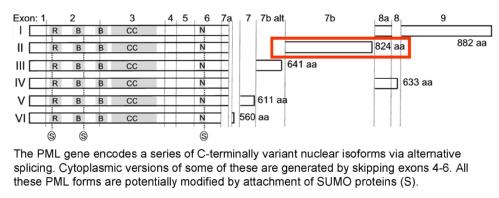
We have shown that the target via which Orf3 disrupts PML-NBs is one specific isoform of PML, PML-II. The interaction involves the self-interaction of Orf3 to form nuclear 'tracks', to which PML is then drawn. All isoforms are recruited because they can interact with one another. The specific target in PML-II is a motif in its unique C-terminal domain.
Currently, we are studying the unique functions of PML-II in host cell biology, and in particular in the innate immune response to infection. SiRNA knock-down specifically of PML-II leads to a profound defect in the induction of interferon responses. This defect arises at the point of assembly of active transcription complexes on genes such as Interferon beta, and interferon-induced genes such as ISG15 and ISG56. Transcription factors that are known to be important for expression of these genes are present but do not associate with the target promoters effectively. The co-activator CBP is also not correctly recruited. We are now investigating how the chromatin at these promoters is altered in the absence of PML-II and have an MRC project to pursue this work, in collaboration with Dr Daniel HebenstreitLink opens in a new window.
PML-NBs have been implicated in the control of infection by many viruses. They are also implicated in cellular stress responses. We are studying the effect of PML-II on the progress of adenovirus infection, either under standard growth conditions or after subjecting cells to mild heat stress. These experiments are testing the idea that stress may materially alter the outcome of infection at the cellular level. We have found that removal of PML-II substantially enhances adenovirus replication. In part, this is the result of the impaired innate immune response that is seen when PML-II is depleted, but a major factor in the increased virus production is the increased level of HSP70 found in cells in which PML-II has been depleted: removal of this induced HSP70 by siRNA knock-down greatly reduces the elevation in viral activity in PML-II knock-down cells.
The temporal cascade of adenovirus gene expression
Adenovirus RNAs are subject to complex patterns of alternative splicing and polyadenylation and a particular focus of interest is to understand how adenovirus proteins can affect RNA processing and RNA transport from the nucleus. One strand of this work has been an interest in the viral E1b 55K protein, deficiency of which leads to a failure to transport and translate late viral mRNA. This protein associates with E4 Orf3, potentially linking our studies of gene expression to our work with PML-NBs (above). The second strand is focused specifically on the adenovirus major late transcription unit (MLTU). This is a very complex gene, encoding close to 20 different proteins from mRNAs produced by alternative splicing and polyadenylation (see map, right).
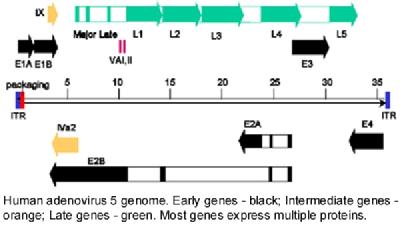
We are studying the role of three viral proteins encoded in the L4 segment of this gene in the proper regulation of its expression and have shown that two of these, 22K and 33K, are crucial up-regulators of full late gene expression. We were surprised to find in 2010 that these proteins are expressed from a dedicated promoter embedded within L4 that is activated transiently at the time of the early-late switch in viral gene expression. More recently, we showed that this promoter is activated by p53, a cellular transcription factor that is activated under various stress conditions. This discovery links the progress of infection to the particular physiological state of the host cell. Most recently, we have shown that TFII-I negatively regulates the L4 promoter, and is overcome by the action of E4 Orf3. As well as giving us important insights into the biology of the virus, which we are exploiting in our vector development studies, we hope that this work will also shed light on important aspects of the host cell gene expression pathway.
Informed by our studies of L4 function in gene expression, we are also working on further developing adenovirus as a vector for gene delivery into cells. Animal and human trials of first generation adenovirus gene therapy vectors have shown that they induce cellular immune responses which then eliminate the transduced cells. The capacity of such vectors for exogenous DNA was also a limitation in some contexts. Vectors with further deletions of viral gene sequences, particularly of the late genes, would be expected to overcome some of these problems. We are developing a number of cell lines which can support growth of adenovirus vectors having deletions in one or more of the late genes of the virus. By removing these late functions, stimulation of antiviral cytotoxic cell responses by the vector in vivo should be reduced and the capacity of the vector for foreign DNA will be increased. This approach links directly from what we have learned about adenovirus gene expression in our fundamental studies of the virus.
